Published paper: The latent resistome
What is the latent resistome? This is a term we coin in a new paper published yesterday in Microbiome. In the paper, we distinguish between the small number antibiotic resistance genes (ARGs) that are established, well-characterized, and available in existing resistance gene databases (what we refer to as “established ARGs”). These are typically ARGs encountered in clinical pathogens and are often already causing problems in human and animal infections. The remaining latently present ARGs, which we denote “latent ARGs”, are less or not at all studied, and are therefore much harder to detect (1). These latent ARGs are typically unknown and generally overlooked in most studies of resistance. They are also seldom accounted for in risk assessments of antibiotic resistance (2-4). This means that our view of the resistome and its diversity is incomplete, which hampers our ability to assess risk for promotion and spread of yet undiscovered resistance determinants.
In our new study, we try to alleviate this issue by analyzing more than 10,000 metagenomic samples. We show that the latent ARGs are more abundant and diverse than established ARGs in all studied environments, including the human- and animal-associated microbiomes. The total pan-resistomes, i.e., all ARGs present in an environment (including the latent ARGs), are heavily dominated by these latent ARGs. In contrast, the core resistome (the ARGs that are commonly encountered) comprise both latent and established ARGs.
In the study, we identified several latent ARGs that were shared between environments or that are already present in human pathogens. These are often located on mobile genetic elements that can be transferred between bacteria. Finally, we also show that wastewater microbiomes have surprisingly large pan- and core-resistomes, which makes this environment a potent high-risk environment for mobilization and promotion of latent ARGs, which may make it into pathogens in the future.
It is also interesting to note that this new study echoes the results of my own study from 2018, showing that soil and water environments contain a high diversity of latent ARGs (or ARGs not found in pathogens as I put it in the 2018 study), despite being almost devoid of established ARGs (5).
This project has been a collaboration with Erik Kristiansson’s research group, and particularly with Juan Inda-Diáz. It has been great fun to work with them and I hope that we will keep this collaboration going into the future! The study can be read in its entirety here.
References
- Inda-Díaz JS, Lund D, Parras-Moltó M, Johnning A, Bengtsson-Palme J, Kristiansson E: Latent antibiotic resistance genes are abundant, diverse, and mobile in human, animal, and environmental microbiomes. Microbiome, 11, 44 (2023). doi: 10.1186/s40168-023-01479-0 [Paper link]
- Martinez JL, Coque TM, Baquero F: What is a resistance gene? Ranking risk in resistomes. Nature Reviews Microbiology 2015, 13:116–123. doi:10.1038/nrmicro3399
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Bengtsson-Palme J: Assessment and management of risks associated with antibiotic resistance in the environment. In: Roig B, Weiss K, Thoreau V (Eds.) Management of Emerging Public Health Issues and Risks: Multidisciplinary Approaches to the Changing Environment, 243–263. Elsevier, UK (2019). doi: 10.1016/B978-0-12-813290-6.00010-X
- Bengtsson-Palme J: The diversity of uncharacterized antibiotic resistance genes can be predicted from known gene variants – but not always. Microbiome, 6, 125 (2018). doi: 10.1186/s40168-018-0508-2
Published paper: Modeling antibiotic resistance gene emergence
Last week, a paper resulting from a collaboration with Stefanie Heß and Viktor Jonsson was published in Environmental Science & Technology. In the paper, we build a quantitative model for the emergence of antibiotic resistance genes in human pathogens and populate it using the few numbers that are available on different processes (bacterial uptake, horizontal gene transfer rates, rate of mobilization of chromosomal genes, etc.) in the literature (1).
In short, we find that in order for the environment to play an important role in the appearance of novel resistance genes in pathogens, there needs to be a substantial flow of bacteria from the environment to the human microbiome. We also find that most likely the majority of resistance genes in human pathogens have very small fitness costs associated with them, if any cost at all.
The model makes three important predictions:
- The majority of ARGs present in pathogens today should have very limited effects on fitness. The model caps the average fitness impact for ARGs currently present in human pathogens between −0.2 and +0.1% per generation. By determining the fitness effects of carrying individual ARGs in their current hosts, this prediction could be experimentally tested.
- The most likely location of ARGs 70 years ago would have been in human-associated bacteria. By tracking ARGs currently present in human pathogens across bacterial genomes, it may be possible to trace the evolutionary history of these genes and thereby identify their likely hosts at the beginning of the antibiotic era, similar to what was done by Stefan Ebmeyer and his colleagues (2). What they found sort-of corroborate the findings of our model and lend support to the idea that most ARGs may not originate in the environment. However, this analysis is complicated by the biased sampling of fully sequenced bacterial genomes, most of which originate from human specimens. That said, the rapid increase in sequencing capacity may make full-scale analysis of ARG origins using genomic data possible in the near future, which would enable testing of this prediction of the model.
- If the origins of ARGs currently circulating in pathogens can be established, the range of reasonable dispersal ability levels from the environment to pathogens narrows dramatically. Similarly, if the rates of mobilization and horizontal transfer of resistance genes could be better determined by experiments, the model would predict the likely origins more precisely. Just establishing a ball-park range for the mobilization rate would dramatically restrict the possible outcomes of the model. Thus, a more precise determination of any of these parameters would enable several more specific predictions by the model.
This paper has a quite interesting backstory, beginning with me having leftover time on a bus ride in Madison (WI), thinking about whether you could quantize the conceptual framework for resistance gene emergence we described in our 2018 review paper in FEMS Reviews Microbiology (3). This spurred the first attempt at such a model, which then led to Stefanie Heß and me applying for support from the Centre for Antibiotic Resistance Research at the University of Gothenburg (CARe) to develop this idea further. We got this support and Stefanie spent a few days with me in Gothenburg developing this idea into a model we could implement in R.
However, at that point we realized we needed more modeling expertise and brought in Viktor Jonsson to make sure the model was robust. From there, it took us about 1.5 years to refine and rerun the model about a million times… By the early spring this year, we had a reasonable model that we could write a manuscript around, and this is what now is published. It’s been an interesting and very nice ride together with Stefanie and Viktor!
References
- Bengtsson-Palme J, Jonsson V, Heß S: What is the role of the environment in the emergence of novel antibiotic resistance genes? A modelling approach. Environmental Science & Technology, Article ASAP (2021). doi: 10.1021/acs.est.1c02977 [Paper link]
- Ebmeyer S, Kristiansson E, Larsson DGJ: A framework for identifying the recent origins of mobile antibiotic resistance genes. Communications Biology 4 (2021). doi: 10.1038/s42003-020-01545-5
- Bengtsson-Palme J, Kristiansson E, Larsson DGJ: Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, 42, 1, 68–80 (2018). doi: 10.1093/femsre/fux053 [Paper link]
March 2021 Pod: Antibiotic resistance evolution
In this episode Microbiology Lab Pod, the team (Johan Bengtsson-Palme, Emil Burman, Anna Abramova, Marcus Wenne, Sebastian Wettersten and Mahbuba Lubna Akter, Shumaila Malik, Emilio Rudbeck and Camille Wuyts) discusses the evolution of antibiotic resistance from different perspectives. We also interview Rémi Gschwind about his work on novel antibiotic resistance genes in the EMBARK program.
The specific papers discussed in the pod (with approximate timings) are as follows:
- 7:45 – EMBARK website: http://antimicrobialresistance.eu
- 26:15 – Seemann, T., 2014. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069. https://doi.org/10.1093/bioinformatics/btu153
- 29:00 – Bengtsson-Palme, J., Larsson, D.G.J., 2015. Antibiotic resistance genes in the environment: prioritizing risks. Nature reviews Microbiology 13, 396. https://doi.org/10.1038/nrmicro3399-c1
- 29:30 – Ebmeyer, S., Kristiansson, E., Larsson, D.G.J., 2021. A framework for identifying the recent origins of mobile antibiotic resistance genes. Communications Biology 4. https://doi.org/10.1038/s42003-020-01545-5
- 54:15 – Gillings, M.R., Stokes, H.W., 2012. Are humans increasing bacterial evolvability? Trends in Ecology & Evolution 27, 346–352. https://doi.org/10.1016/j.tree.2012.02.006
- 55:15 – Woods, L.C., et al., 2020. Horizontal gene transfer potentiates adaptation by reducing selective constraints on the spread of genetic variation. Proc Natl Acad Sci USA 117, 26868–26875. https://doi.org/10.1073/pnas.2005331117
- 76:15 – Card, K.J., Thomas, M.D., Graves, J.L., Barrick, J.E., Lenski, R.E., 2021. Genomic evolution of antibiotic resistance is contingent on genetic background following a long-term experiment with Escherichia coli. Proc Natl Acad Sci USA 118, e2016886118. https://doi.org/10.1073/pnas.2016886118
The podcast was recorded on March 18, 2021. If you want to reach out to us with comments, suggestions, or other feedback, please send an e-mail to podcast at microbiology dot se or contact @bengtssonpalme via Twitter. The music that can be heard on the pod is composed by Johan Bengtsson-Palme and is taken from the album Cafe Phonocratique.
Podcast: Play in new window | Download
Subscribe: RSS
September 2020 Pod: All antibiotic resistance
This is the fifth episode of the Microbiology Lab Pod and has been lying around on my computer almost finished for way too long. It was recorded on September 23, and the bigger-than-ever-before crew (Johan Bengtsson-Palme, Emil Burman, Haveela Kunche, Anna Abramova, Marcus Wenne, Sebastian Wettersten and Mahbuba Lubna Akter) is joined by Fanny Berglund to discuss computational discovery of novel resistance genes. We also discuss antibiotic resistance mechanisms, particularly in Pseudomonas aeruginosa.
The specific papers discussed in the pod (with approximate timings) are as follows:
- 5:30 – Berglund, F., Johnning, A., Larsson, D.G.J., Kristiansson, E., 2020. An updated phylogeny of the metallo-b-lactamases. Journal of Antimicrobial Chemotherapy 7. https://doi.org/10.1093/jac/dkaa392
- 5:45 – Berglund, F., Österlund, T., Boulund, F., Marathe, N.P., Larsson, D.G.J., Kristiansson, E., 2019. Identification and reconstruction of novel antibiotic resistance genes from metagenomes. Microbiome 7, 52. https://doi.org/10.1186/s40168-019-0670-1
- 6:00 – Berglund, F., Marathe, N.P., Österlund, T., Bengtsson-Palme, J., Kotsakis, S., Flach, C.-F., Larsson, D.G.J., Kristiansson, E., 2017. Identification of 76 novel B1 metallo-β-lactamases through large-scale screening of genomic and metagenomic data. Microbiome 5, i29. https://doi.org/10.1186/s40168-017-0353-8
- 6:15 – Boulund, F., Berglund, F., Flach, C.-F., Bengtsson-Palme, J., Marathe, N.P., Larsson, D.G.J., Kristiansson, E., 2017. Computational discovery and functional validation of novel fluoroquinolone resistance genes in public metagenomic data sets. BMC Genomics 18, 438. https://doi.org/10.1186/s12864-017-4064-0
- 37:15 – Crippen, C.S., Jr., M.J.R., Sanchez, S., Szymanski, C.M., 2020. Multidrug Resistant Acinetobacter Isolates Release Resistance Determinants Through Contact-Dependent Killing and Bacteriophage Lysis. Frontiers in Microbiology 11. https://doi.org/10.3389/fmicb.2020.01918
- 52:15 – Leonard, A.F.C., Zhang, L., Balfour, A.J., Garside, R., Hawkey, P.M., Murray, A.K., Ukoumunne, O.C., Gaze, W.H., 2018. Exposure to and colonisation by antibiotic-resistant E. coli in UK coastal water users: Environmental surveillance, exposure assessment, and epidemiological study (Beach Bum Survey). Environment International 114, 326–333. https://doi.org/10.1016/j.envint.2017.11.003
- 53:30 – Bengtsson-Palme, J., Kristiansson, E., Larsson, D.G.J., 2018. Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews 42, 25. https://doi.org/10.1093/femsre/fux053
- 54:30 – Leonard, A.F.C., Zhang, L., Balfour, A.J., Garside, R., Gaze, W.H., 2015. Human recreational exposure to antibiotic resistant bacteria in coastal bathing waters. Environment International 82, 92–100. https://doi.org/10.1016/j.envint.2015.02.013
- 55:30 – Ahmed, M.N., Abdelsamad, A., Wassermann, T., et al., 2020. The evolutionary trajectories of P. aeruginosa in biofilm and planktonic growth modes exposed to ciprofloxacin: beyond selection of antibiotic resistance. npj Biofilms and Microbiomes 6. https://doi.org/10.1038/s41522-020-00138-8
- 69:30 – Rezzoagli, C., Archetti, M., Mignot, I., Baumgartner, M., Kümmerli, R., 2020. Combining antibiotics with antivirulence compounds can have synergistic effects and reverse selection for antibiotic resistance in Pseudomonas aeruginosa. PLOS Biology 18, e3000805. https://doi.org/10.1371/journal.pbio.3000805
- 79:45 – Allen, R.C., Popat, R., Diggle, S.P., Brown, S.P., 2014. Targeting virulence: can we make evolution-proof drugs? Nature reviews Microbiology 12, 300–308. https://doi.org/10.1038/nrmicro3232
- 80:45 – Köhler, T., Perron, G.G., Buckling, A., van Delden, C., 2010. Quorum Sensing Inhibition Selects for Virulence and Cooperation in Pseudomonas aeruginosa. PLoS Pathogens 6, e1000883. https://doi.org/10.1371/journal.ppat.1000883
The podcast was recorded on September 23, 2020. If you want to reach out to us with comments, suggestions or other feedback, please send an e-mail to podcast at microbiology dot se or contact @bengtssonpalme via Twitter. The music that can be heard on the pod is composed by Johan Bengtsson-Palme and is taken from the album Cafe Phonocratique.
Podcast: Play in new window | Download
Subscribe: RSS
May 2020 Pod: Discovering novel resistance genes and how bacteria become virulent
In the second episode of Microbiology Lab Pod, a crew consisting of Johan Bengtsson-Palme, Emil Burman, Haveela Kunche and Anna Abramova discusses how to identify novel resistance genes with our special guest Marlies Böhm. We also talk about bacterial virulence: how do bacteria become virulent, how do virulence relate to competition, how do bacteria evade the immune system and can we attenuate virulence using fatty acids?
The specific papers discussed in the pod (with approximate timings) are as follows:
- 7:15 – Böhm, M.-E., Razavi, M., Flach, C.-F., Larsson, D.G.J., 2020a. A Novel, Integron-Regulated, Class C β-Lactamase. Antibiotics 9, 123. https://doi.org/10.3390/antibiotics9030123
- 7:15 – Böhm, M.-E., Razavi, M., Marathe, N.P., Flach, C.-F., Larsson, D.G.J., 2020b. Discovery of a novel integron-borne aminoglycoside resistance gene present in clinical pathogens by screening environmental bacterial communities. Microbiome 8. https://doi.org/10.1186/s40168-020-00814-z
- 9:15 – Makowska, N., et al., 2020. Occurrence of integrons and antibiotic resistance genes in cryoconite and ice of Svalbard, Greenland, and the Caucasus glaciers. Science of The Total Environment 716, 137022. https://doi.org/10.1016/j.scitotenv.2020.137022
- 20:45 – Marathe, N.P., et al., 2019. Scandinavium goeteborgense gen. nov., sp. nov., a New Member of the Family Enterobacteriaceae Isolated From a Wound Infection, Carries a Novel Quinolone Resistance Gene Variant. Frontiers in Microbiology 10. https://doi.org/10.3389/fmicb.2019.02511
- 33:45 – Kaito, C., Yoshikai, H., Wakamatsu, A., Miyashita, A., Matsumoto, Y., Fujiyuki, T., Kato, M., Ogura, Y., Hayashi, T., Isogai, T., Sekimizu, K., 2020. Non-pathogenic Escherichia coli acquires virulence by mutating a growth-essential LPS transporter. PLOS Pathogens 16, e1008469. https://doi.org/10.1371/journal.ppat.1008469
- 43:45 – Lories, B., Roberfroid, S., Dieltjens, L., De Coster, D., Foster, K.R., Steenackers, H.P., 2020. Biofilm Bacteria Use Stress Responses to Detect and Respond to Competitors. Current Biology 30, 1231-1244.e4. https://doi.org/10.1016/j.cub.2020.01.065
- 45:45 – Lozano, G.L., Bravo, J.I., Garavito Diago, M.F., Park, H.B., Hurley, A., Peterson, S.B., Stabb, E.V., Crawford, J.M., Broderick, N.A., Handelsman, J., 2019. Introducing THOR, a Model Microbiome for Genetic Dissection of Community Behavior. mBio 10. https://doi.org/10.1128/mBio.02846-18
- 55:45 – Kumar, P., Lee, J.-H., Beyenal, H., Lee, J., 2020. Fatty Acids as Antibiofilm and Antivirulence Agents. Trends in Microbiology. https://doi.org/10.1016/j.tim.2020.03.014
- 60:15 – Gullberg, E., Cao, S., Berg, O.G., Ilbäck, C., Sandegren, L., Hughes, D., Andersson, D.I., 2011. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathogens 7, e1002158. https://doi.org/10.1371/journal.ppat.1002158
- 61:15 – Larsson, D.G.J., 2018. Risks of using the natural defence of commensal bacteria as antibiotics call for research and regulation. International Journal of Antimicrobial Agents 51, 277–278. https://doi.org/10.1016/j.ijantimicag.2017.12.018
- 65:15 – Lone, A.G., Bankhead, T., 2020. The Borrelia burgdorferi VlsE Lipoprotein Prevents Antibody Binding to an Arthritis-Related Surface Antigen. Cell Reports 30, 3663-3670.e5. https://doi.org/10.1016/j.celrep.2020.02.081
The podcast was recorded on May 7, 2020. If you want to reach out to us with comments, suggestions or other feedback, please send an e-mail to podcast at microbiology dot se or contact @bengtssonpalme via Twitter. The music that can be heard on the pod is composed by Johan Bengtsson-Palme and is taken from the album Cafe Phonocratique.
Podcast: Play in new window | Download
Subscribe: RSS
Conferences and a PhD position
Here’s some updates on my Spring schedule.
On March 19, I will be presenting the EMBARK program and what we aim to achieve at a conference organised by the Swedish Medical Products Agency called NordicMappingAMR. The event will feature an overview of existing monitoring of antibiotics and antibiotic resistant bacteria in the environment. The conference aims to present the results from this survey, to listen to experts in the field and to discuss possible progress. It takes place in Uppsala. For any further questions, contact Kia Salin at NordicMappingAMR@lakemedelsverket.se
Then on May 18 to 20 I will participate in the 7th Microbiome & Probiotics R&D and Business Collaboration Forum in Rotterdam. This industry/academia cross-over event focuses on cutting-edge microbiome and probiotics research, and challenges and opportunities in moving research towards commercialisation. I will talk on the work we do on deciphering genetic mechanisms behind microbial interactions in microbiomes on May 20.
And finally, I also want to bring the attention to that my collaborator Erik Kristiansson has an open PhD position in his lab. The position is funded by the Environmental Dimensions of Antibiotic Resistance (EDAR) research project, aiming to describe the environmental role in the development and promotion of antibiotic resistance. The focus of the PhD position will be on analysis of large-scale data, with special emphasis on the identification of new forms of resistance genes. The project also includes phylogenetic analysis and development of methods for assessment of gene evolution. More info can be found here.
Published paper: Knowledge gaps for environmental antibiotic resistance
The outcomes from a workshop arranged by JPIAMR, the Swedish Research Council (VR) and CARe were just published as a short review paper in Environment International. In the paper, which was mostly moved forward by Prof. Joakim Larsson at CARe, we describe four major areas of knowledge gaps in the realm of environmental antibiotic resistance (1). We then highlight several important sub-questions within these areas. The broad areas we define are:
- What are the relative contributions of different sources of antibiotics and antibiotic resistant bacteria into the environment?
- What is the role of the environment as affected by anthropogenic inputs (e.g. pollution and other activities) on the evolution (mobilization, selection, transfer, persistence etc.) of antibiotic resistance?
- How significant is the exposure of humans to antibiotic resistant bacteria via different environmental routes, and what is the impact on human health?
- What technological, social, economic and behavioral interventions are effective to mitigate the emergence and spread of antibiotic resistance via the environment?
Although much has been written on the topic before (e.g. 2-12), I think it is unique that we collect and explicitly point out areas in which we are lacking important knowledge to build accurate risk models and devise appropriate intervention strategies. The workshop was held in Gothenburg on the 27–28th of September 2017. The workshop leaders Joakim Larsson, Ana-Maria de Roda Husman and Ramanan Laxminarayan were each responsible for moderating a breakout group, and every breakout group was tasked to deal with knowledge gaps related to either evolution, transmission or interventions. The reports of the breakout groups were then discussed among all participants to clarify and structure the areas where more research is needed, which boiled down to the four overarching critical knowledge gaps described in the paper (1).
This is a short paper, and I think everyone with an interest in environmental antibiotic resistance should read it and reflect over its content (because, we may of course have overlooked some important aspect). You can find the paper here.
References
- Larsson DGJ, Andremont A, Bengtsson-Palme J, Brandt KK, de Roda Husman AM, Fagerstedt P, Fick J, Flach C-F, Gaze WH, Kuroda M, Kvint K, Laxminarayan R, Manaia CM, Nielsen KM, Ploy M-C, Segovia C, Simonet P, Smalla K, Snape J, Topp E, van Hengel A, Verner-Jeffreys DW, Virta MPJ, Wellington EM, Wernersson A-S: Critical knowledge gaps and research needs related to the environmental dimensions of antibiotic resistance. Environment International, 117, 132–138 (2018). doi: 10.1016/j.envint.2018.04.041
- Bengtsson-Palme J, Kristiansson E, Larsson DGJ: Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, 42, 1, 68–80 (2018). doi: 10.1093/femsre/fux053
- Martinez JL, Coque TM, Baquero F: What is a resistance gene? Ranking risk in resistomes. Nature Reviews Microbiology 2015, 13:116–123. doi:10.1038/nrmicro3399
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Ashbolt NJ, Amézquita A, Backhaus T, Borriello P, Brandt KK, Collignon P, et al.: Human Health Risk Assessment (HHRA) for Environmental Development and Transfer of Antibiotic Resistance. Environmental Health Perspectives, 121, 993–1001 (2013)
- Pruden A, Larsson DGJ, Amézquita A, Collignon P, Brandt KK, Graham DW, et al.: Management options for reducing the release of antibiotics and antibiotic resistance genes to the environment. Environmental Health Perspectives, 121, 878–85 (2013).
- Gillings MR: Evolutionary consequences of antibiotic use for the resistome, mobilome and microbial pangenome. Frontiers in Microbiology, 4, 4 (2013).
- Baquero F, Alvarez-Ortega C, Martinez JL: Ecology and evolution of antibiotic resistance. Environmental Microbiology Reports, 1, 469–476 (2009).
- Baquero F, Tedim AP, Coque TM: Antibiotic resistance shaping multi-level population biology of bacteria. Frontiers in Microbiology, 4, 15 (2013).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Hiltunen T, Virta M, Laine A-L: Antibiotic resistance in the wild: an eco-evolutionary perspective. Philosophical Transactions of the Royal Society B: Biological Sciences, 372 (2017) doi: 10.1098/rstb.2016.0039.
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
Published paper: Environmental factors leading to resistance
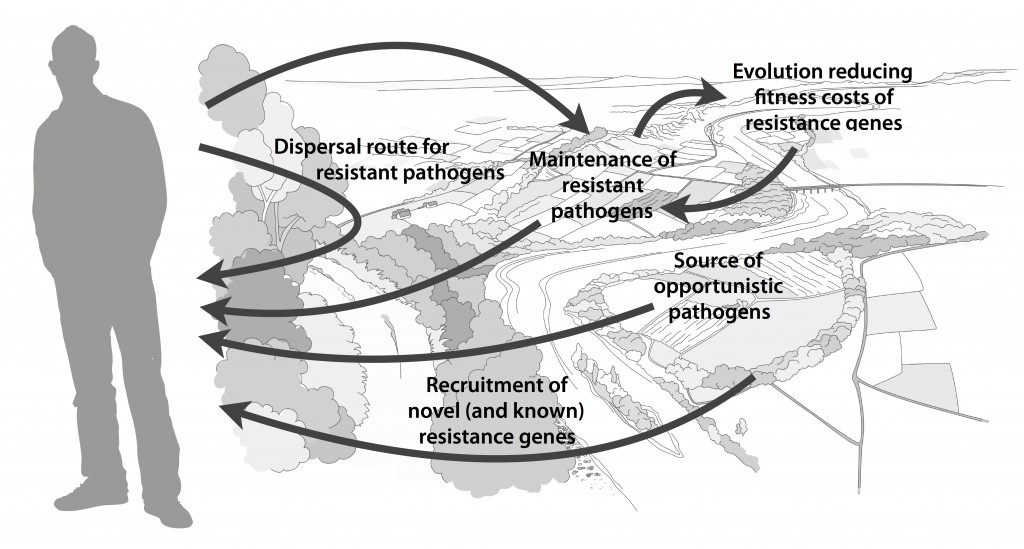
Myself, Joakim Larsson and Erik Kristiansson have written a review on the environmental factors that influence development and spread of antibiotic resistance, which was published today in FEMS Microbiology Reviews. The review (1) builds on thoughts developed in the latter parts of my PhD thesis (2), and seeks to provide a synthesis knowledge gained from different subfields towards the current understanding of evolutionary and ecological processes leading to clinical appearance of resistance genes, as well as the important environmental dispersal barriers preventing spread of resistant pathogens.
We postulate that emergence of novel resistance factors and mobilization of resistance genes are likely to occur continuously in the environment. However, the great majority of such genetic events are unlikely to lead to establishment of novel resistance factors in bacterial populations, unless there is a selection pressure for maintaining them or their fitness costs are negligible. To enable measures to prevent resistance development in the environment, it is therefore critical to investigate under what conditions and to what extent environmental selection for resistance takes place. Selection for resistance is likely less important for the dissemination of resistant bacteria, but will ultimately depend on how well the species or strain in question thrives in the external environment. Metacommunity theory (3,4) suggests that dispersal ability is central to this process, and therefore opportunistic pathogens with their main habitat in the environment may play an important role in the exchange of resistance factors between humans and the environment. Understanding the dispersal barriers hindering this exchange is not only key to evaluate risks, but also to prevent resistant pathogens, as well as novel resistance genes, from reaching humans.
Towards the end of the paper, we suggest certain environments that seem to be more important from a risk management perspective. We also discuss additional problems linked to the development of antibiotic resistance, such as increased evolvability of bacterial genomes (5) and which other types of genes that may be mobilized in the future, should the development continue (1,6). In this review, we also further develop thoughts on the relative risks of re-recruiting and spreading well-known resistance factors already circulating in pathogens, versus recruitment of completely novel resistance genes from environmental bacteria (7). While the latter case is likely to be very rare, and thus almost impossible to quantify the risks for, the consequences of such (potentially one-time) events can be dire.
I personally think that this is one of the best though-through pieces I have ever written, and since it is open access and (in my biased opinion) written in a fairly accessible way, I recommend everyone to read it. It builds on the ecological theories for resistance ecology developed by, among others, Fernando Baquero and José Martinez (8-13). Over the last year, it has been stressed several times at meetings (e.g. at the EDAR conferences in August) that there is a need to develop an ecological framework for antibiotic resistance genes. I think this paper could be one of the foundational pillars on such an endeavor and look forward to see how it will fit into the growing literature on the subject!
References
- Bengtsson-Palme J, Kristiansson E, Larsson DGJ: Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, accepted manuscript (2017). doi: 10.1093/femsre/fux053
- Bengtsson-Palme J: Antibiotic resistance in the environment: a contribution from metagenomic studies. Doctoral thesis (medicine), Department of Infectious Diseases, Institute of Biomedicine, Sahlgrenska Academy, University of Gothenburg, 2016. [Link]
- Bengtsson J: Applied (meta)community ecology: diversity and ecosystem services at the intersection of local and regional processes. In: Verhoef HA, Morin PJ (eds.). Community Ecology: Processes, Models, and Applications. Oxford: Oxford University Press, 115–130 (2009).
- Leibold M, Norberg J: Biodiversity in metacommunities: Plankton as complex adaptive systems? Limnology and Oceanography, 1278–1289 (2004).
- Gillings MR, Stokes HW: Are humans increasing bacterial evolvability? Trends in Ecology and Evolution, 27, 346–352 (2012).
- Gillings MR: Evolutionary consequences of antibiotic use for the resistome, mobilome and microbial pangenome. Frontiers in Microbiology, 4, 4 (2013).
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Baquero F, Alvarez-Ortega C, Martinez JL: Ecology and evolution of antibiotic resistance. Environmental Microbiology Reports, 1, 469–476 (2009).
- Baquero F, Tedim AP, Coque TM: Antibiotic resistance shaping multi-level population biology of bacteria. Frontiers in Microbiology, 4, 15 (2013).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Hiltunen T, Virta M, Laine A-L: Antibiotic resistance in the wild: an eco-evolutionary perspective. Philosophical Transactions of the Royal Society B: Biological Sciences, 372 (2017) doi: 10.1098/rstb.2016.0039.
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
- Salyers AA, Amábile-Cuevas CF: Why are antibiotic resistance genes so resistant to elimination? Antimicrobial Agents and Chemotherapy, 41, 2321–2325 (1997).
Published paper: 76 new metallo-beta-lactamases
Today, Microbiome put online a paper lead-authored by my colleague Fanny Berglund – one of Erik Kristiansson‘s brilliant PhD students – in which we identify 76 novel metallo-ß-lactamases (1). This feat was made possible because of a new computational method designed by Fanny, which uses a hidden Markov model based on known B1 metallo-ß-lactamases. We analyzed over 10,000 bacterial genomes and plasmids and over 5 terabases of metagenomic data and could thereby predict 76 novel genes. These genes clustered into 59 new families of metallo-β-lactamases (given a 70% identity threshold). We also verified the functionality of 21 of these genes experimentally, and found that 18 were able to hydrolyze imipenem when inserted into Escherichia coli. Two of the novel genes contained atypical zinc-binding motifs in their active sites. Finally, we show that the B1 metallo-β-lactamases can be divided into five major groups based on their phylogenetic origin. It seems that nearly all of the previously characterized mobile B1 β-lactamases we identify in this study were likely to have originated from chromosomal genes present in species within the Proteobacteria, particularly Shewanella spp.
This study more than doubles the number of known B1 metallo-β-lactamases. As with the study by Boulund et al. (2) which we published last month on computational discovery of novel fluoroquinolone resistance genes (which used a very similar approach but on a completely different type of genes), this study also supports the hypothesis that environmental bacterial communities act as sources of uncharacterized antibiotic resistance genes (3-7). Fanny have done a fantastic job on this paper, and I highly recommend reading it in its entirety (it’s open access so you have virtually no excuse not to). It can be found here.
References
- Berglund F, Marathe NP, Österlund T, Bengtsson-Palme J, Kotsakis S, Flach C-F, Larsson DGJ, Kristiansson E: Identification of 76 novel B1 metallo-β-lactamases through large-scale screening of genomic and metagenomic data. Microbiome, 5, 134 (2017). doi: 10.1186/s40168-017-0353-8
- Boulund F, Berglund F, Flach C-F, Bengtsson-Palme J, Marathe NP, Larsson DGJ, Kristiansson E: Computational discovery and functional validation of novel fluoroquinolone resistance genes in public metagenomic data sets. BMC Genomics, 18, 682 (2017). doi: 10.1186/s12864-017-4064-0
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Allen HK, Donato J, Wang HH et al.: Call of the wild: antibiotic resistance genes in natural environments. Nature Reviews Microbiology, 8, 251–259 (2010).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
- Finley RL, Collignon P, Larsson DGJ et al.: The scourge of antibiotic resistance: the important role of the environment. Clinical Infectious Diseases, 57, 704–710 (2013).
Published paper: Computational discovery of novel qnr genes
BMC Genomics today published a paper first-authored by my long-time colleague Fredrik Boulund, which describes a computational screen of genomes and metagenomes for novel qnr fluoroquinolone resistance genes (1). The study makes use of Fredrik’s well-designed and updated qnr-prediction pipeline, but in contrast to his previous publication based on the pipeline from 2012 (2), we here study a 20-fold larger dataset of almost 13 terabases of sequence data. Based on this data, the pipeline predicted 611 putative qnr genes, including all previously described plasmid-mediated qnr gene families. 20 of the predicted genes were previously undescribed, and of these nine were selected for experimental validation. Six of those tested genes improved the survivability under ciprofloxacin exposure when expressed in Escherichia coli. The study shows that qnr genes are almost ubiquitous in environmental microbial communities. This study also lends further credibility to the hypothesis that environmental bacterial communities can act as sources of previously uncharacterized antibiotic resistance genes (3-7). The study can be read in its entirety here.
References
- Boulund F, Berglund F, Flach C-F, Bengtsson-Palme J, Marathe NP, Larsson DGJ, Kristiansson E: Computational discovery and functional validation of novel fluoroquinolone resistance genes in public metagenomic data sets. BMC Genomics, 18, 682 (2017). doi: 10.1186/s12864-017-4064-0
- Boulund F, Johnning A, Pereira MB, Larsson DGJ, Kristiansson E: A novel method to discover fluoroquinolone antibiotic resistance (qnr) genes in fragmented nucleotide sequences. BMC Genomics, 13, 695 (2012). doi: 10.1186/1471-2164-13-695
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Allen HK, Donato J, Wang HH et al.: Call of the wild: antibiotic resistance genes in natural environments. Nature Reviews Microbiology, 8, 251–259 (2010).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
- Finley RL, Collignon P, Larsson DGJ et al.: The scourge of antibiotic resistance: the important role of the environment. Clinical Infectious Diseases, 57, 704–710 (2013).