Published book chapter: Strategies for metagenomic analysis
Last summer, I was approached by Muniyandi Nagarajan to write a book chapter for a book on metagenomics. The book was published earlier this month, and is now available online (1). I have to admit that I have not yet read the entire book, but my own chapter deals with selecting the right tools for metagenomic analysis, and discusses different strategies to perform taxonomic classification, functional analysis, metagenomic assembly, and statistical comparisons between metagenomes (2). The chapter also considers the pros and cons of automated computational “pipelines” for analysis of metagenomic data. While I do not point to a specific set of software that obviously perform better in all situations, I do highlight some analysis strategies that clearly should be avoided. The chapter also suggests a few among the set of robust and well-functioning software tools that, in my opinion, should be used for metagenomic analyses. To some degree, this paper overlaps with the review paper we wrote on using metagenomics to analyze antibiotic resistance genes in various environments, published earlier this year (3), but the discussion in the book chapter is far more general. I imagine that the book chapter could be used, for example, in teaching metagenomics to students in bioinformatics (that’s at least a use I envision myself). Finally, apart from my own chapter, I can also highly recommend the chapter by Boulund et al. on statistical considerations for metagenomic data analysis (4). The book is available to buy from here, and the chapter can be read here.
References
- Nagarajan M (Ed.) Metagenomics: Perspectives, Methods, and Applications. ISBN: 9780081022689. Academic Press, Elsevier, USA (2018). doi: 10.1016/B978-0-08-102268-9 [Link]
- Bengtsson-Palme J: Strategies for Taxonomic and Functional Annotation of Metagenomes. In: Nagarajan M (Ed.) Metagenomics: Perspectives, Methods, and Applications, 55–79. Academic Press, Elsevier, USA (2018). doi: 10.1016/B978-0-08-102268-9.00003-3 [Link]
- Bengtsson-Palme J, Larsson DGJ, Kristiansson E: Using metagenomics to investigate human and environmental resistomes. Journal of Antimicrobial Chemotherapy, 72, 2690–2703 (2017). doi: 10.1093/jac/dkx199 [Paper link]
- Boulund F, Pereira MB, Jonsson V, Kristiansson E: Computational and Statistical Considerations in the Analysis of Metagenomic Data. In: Nagarajan M (Ed.) Metagenomics: Perspectives, Methods, and Applications, 81–102. Academic Press,, Elsevier, USA (2018). doi: 10.1016/B978-0-08-102268-9.00004-5 [Link]
Published paper: 76 new metallo-beta-lactamases
Today, Microbiome put online a paper lead-authored by my colleague Fanny Berglund – one of Erik Kristiansson‘s brilliant PhD students – in which we identify 76 novel metallo-ß-lactamases (1). This feat was made possible because of a new computational method designed by Fanny, which uses a hidden Markov model based on known B1 metallo-ß-lactamases. We analyzed over 10,000 bacterial genomes and plasmids and over 5 terabases of metagenomic data and could thereby predict 76 novel genes. These genes clustered into 59 new families of metallo-β-lactamases (given a 70% identity threshold). We also verified the functionality of 21 of these genes experimentally, and found that 18 were able to hydrolyze imipenem when inserted into Escherichia coli. Two of the novel genes contained atypical zinc-binding motifs in their active sites. Finally, we show that the B1 metallo-β-lactamases can be divided into five major groups based on their phylogenetic origin. It seems that nearly all of the previously characterized mobile B1 β-lactamases we identify in this study were likely to have originated from chromosomal genes present in species within the Proteobacteria, particularly Shewanella spp.
This study more than doubles the number of known B1 metallo-β-lactamases. As with the study by Boulund et al. (2) which we published last month on computational discovery of novel fluoroquinolone resistance genes (which used a very similar approach but on a completely different type of genes), this study also supports the hypothesis that environmental bacterial communities act as sources of uncharacterized antibiotic resistance genes (3-7). Fanny have done a fantastic job on this paper, and I highly recommend reading it in its entirety (it’s open access so you have virtually no excuse not to). It can be found here.
References
- Berglund F, Marathe NP, Österlund T, Bengtsson-Palme J, Kotsakis S, Flach C-F, Larsson DGJ, Kristiansson E: Identification of 76 novel B1 metallo-β-lactamases through large-scale screening of genomic and metagenomic data. Microbiome, 5, 134 (2017). doi: 10.1186/s40168-017-0353-8
- Boulund F, Berglund F, Flach C-F, Bengtsson-Palme J, Marathe NP, Larsson DGJ, Kristiansson E: Computational discovery and functional validation of novel fluoroquinolone resistance genes in public metagenomic data sets. BMC Genomics, 18, 682 (2017). doi: 10.1186/s12864-017-4064-0
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Allen HK, Donato J, Wang HH et al.: Call of the wild: antibiotic resistance genes in natural environments. Nature Reviews Microbiology, 8, 251–259 (2010).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
- Finley RL, Collignon P, Larsson DGJ et al.: The scourge of antibiotic resistance: the important role of the environment. Clinical Infectious Diseases, 57, 704–710 (2013).
ITSx in Bioconda
Mattias de Hollander at the Netherlands Institute of Ecology kindly informed me that they recently added the ITSx 1.1b version to the Bioconda package manager. This will make it easy for Conda users to install ITSx automatically into their systems and pipelines and also for others who are using conda. The Bioconda version can be found here. I would like to thank Mattias for this initiative and hope that the Bioconda version of ITSx will find much use!
Published paper: Computational discovery of novel qnr genes
BMC Genomics today published a paper first-authored by my long-time colleague Fredrik Boulund, which describes a computational screen of genomes and metagenomes for novel qnr fluoroquinolone resistance genes (1). The study makes use of Fredrik’s well-designed and updated qnr-prediction pipeline, but in contrast to his previous publication based on the pipeline from 2012 (2), we here study a 20-fold larger dataset of almost 13 terabases of sequence data. Based on this data, the pipeline predicted 611 putative qnr genes, including all previously described plasmid-mediated qnr gene families. 20 of the predicted genes were previously undescribed, and of these nine were selected for experimental validation. Six of those tested genes improved the survivability under ciprofloxacin exposure when expressed in Escherichia coli. The study shows that qnr genes are almost ubiquitous in environmental microbial communities. This study also lends further credibility to the hypothesis that environmental bacterial communities can act as sources of previously uncharacterized antibiotic resistance genes (3-7). The study can be read in its entirety here.
References
- Boulund F, Berglund F, Flach C-F, Bengtsson-Palme J, Marathe NP, Larsson DGJ, Kristiansson E: Computational discovery and functional validation of novel fluoroquinolone resistance genes in public metagenomic data sets. BMC Genomics, 18, 682 (2017). doi: 10.1186/s12864-017-4064-0
- Boulund F, Johnning A, Pereira MB, Larsson DGJ, Kristiansson E: A novel method to discover fluoroquinolone antibiotic resistance (qnr) genes in fragmented nucleotide sequences. BMC Genomics, 13, 695 (2012). doi: 10.1186/1471-2164-13-695
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Allen HK, Donato J, Wang HH et al.: Call of the wild: antibiotic resistance genes in natural environments. Nature Reviews Microbiology, 8, 251–259 (2010).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
- Finley RL, Collignon P, Larsson DGJ et al.: The scourge of antibiotic resistance: the important role of the environment. Clinical Infectious Diseases, 57, 704–710 (2013).
Published paper: The Calanus glacialis mitogenome
Mitochondrial DNA Part B today published a mitochondrial genome announcement paper (1) in which I was involved in doing the assemblies and annotating them. The paper describes the mitogenome of Calanus glacialis, a marine planktonic copepod, which is a keystone species in the Arctic Ocean. The mitogenome is 20,674 bp long, and includes 13 protein-coding genes, 2 rRNA genes and 22 tRNA genes. While this is of course note a huge paper, we believe that this new resource will be of interest in understanding the structure and dynamics of C. glacialis populations. The main work in this paper has been carried out by Marvin Choquet at Nord University in Bodø, Norway. So hats off to him for great work, thanks Marvin! The paper can be read here.
Reference
- Choquet M, Alves Monteiro HJ, Bengtsson-Palme J, Hoarau G: The complete mitochondrial genome of the copepod Calanus glacialis. Mitochondrial DNA Part B, 2, 2, 506–507 (2017). doi: 10.1080/23802359.2017.1361357 [Paper link]
Published paper: Helicobacter pylori vs. the viable gastric microbiota
I have just returned from a week of vacation in Sicily (almost without internet access), so I am a tad late to this news, but earlier this week Infection and Immunity published our paper on the Helicobacter pylori transcriptome in gastric infection (and early stages of carcinogenesis), and how that relates to the transcriptionally active microbiota in the stomach environment (1). This paper has been long in the making (an earlier version of it was included in Kaisa Thorell’s PhD thesis (2)), but some late additional analyses did substantially strengthen our confidence in the suggestions we got from the original data.
In the paper (1) we use metatranscriptomic RNAseq to investigate the composition of the viable microbial community, and at the same time study H. pylori gene expression in stomach biopsies. The biopsies were sampled from individuals with different degrees of H. pylori infection and/or pre-malignant tissue changes. We found that H. pylori completely dominates the microbiota in infected individuals, but (somewhat surprisingly) also in the majority of individuals classified as H. pylori uninfected using traditional methods. This confirms previous reports that have detected minute quantities of H. pylori also in presumably uninfected individuals (3-6), and raises the question of how large part of the human population (if any) that is truly not infected/colonized by H. pylori. The abundance of H. pylori was correlated with the abundance of Campylobacter, Deinococcus, and Sulfurospirillum. It is unclear, however, if these genera only share the same habitat preferences as Helicobacter, or if they are specifically promoted by the presence of H. pylori (or tissue changes induced by it). We also found that genes involved in pH regulation and nickel transport were highly expressed in H. pylori, regardless of disease stage. As far as we know, this study is the first to use metatranscriptomics to study the viable microbiota of the human stomach, and we think that this is a promising approach for future studies on pathogen-microbiota interactions. The paper (in unedited format) can be read here.
References
- Thorell K, Bengtsson-Palme J, Liu OH, Gonzales RVP, Nookaew I, Rabeneck L, Paszat L, Graham DY, Nielsen J, Lundin SB, Sjöling Å: In vivo analysis of the viable microbiota and Helicobacter pylori transcriptome in gastric infection and early stages of carcinogenesis. Infection and Immunity, accepted manuscript (2017). doi: 10.1128/IAI.00031-17 [Paper link]
- Thorell K: Multi-level characterization of host and pathogen in Helicobacter pylori-associated gastric carcinogenesis. Doctoral thesis, Institute of Biomedicine, Sahlgrenska Academy, University of Gothenburg (2014). [Link]
- Bik EM, Eckburg PB, Gill SR, Nelson KE, Purdom EA, Francois F, Perez-Perez G, Blaser MJ, Reman DA: Molecular analysis of the bacterial microbiota in the human stomach. PNAS, 103:732-737 (2006).
- Dicksved J, Lindberg M, Rosenquist M, Enroth H, Jansson JK, Engstrand L: Molecular characterization of the stomach microbiota in patients with gastric cancer and in controls. Journal of Medical Microbiology, 58:509-516 (2009).
- Maldonado-Contreras A, Goldfarb KC, Godoy-Vitorino F, Karaoz U, Contreras M, Blaser MJ, Brodie EL, Dominguez-Bello MG: Structure of the human gastric bacterial community in relation to Helicobacter pylori status. ISME Journal, 5:574-579 (2011).
- Li TH, Qin Y, Sham PC, Lau KS, Chu KM, Leung WK: Alterations in Gastric Microbiota After H. Pylori Eradication and in Different Histological Stages of Gastric Carcinogenesis. Scientific Reports, 7:44935 (2017).
Major ITSx update (beta version)
Today, I am very happy to announce that after years in the making and months in testing, the next generation of ITSx, version 1.1, is ready to step into the public light and scrutiny. I have today uploaded a public beta version of the ITSx 1.1 release, which I encourage everyone that have enjoyed using ITSx to try out.
The 1.1 release of ITSx includes a wide range of new feature, including:
- A 2-10x performance increase (depending on the dataset), since ITSx now utilizes hmmsearch instead of hmmscan to detect the ITS regions and distributes the CPU cores better
- Improved ITS detection among fungi and chlorophyta, by addition of new HMM-profiles
- The HMM profile format for ITSx has been updated to HMMER3/f (thus ITSx now requires HMMER version 3.1 or later)
- Better handling of interrupted HMMER searches
- Added the
--require_anchoroption to only include sequences where the complete anchor is found in the output - Added the possibility for partial sequence output for the SSU, LSU and 5.8S regions
- Fixed a bug causing problems when reading sequence data from standard input
A lot of the code has changed in this version, which means that there might still be bugs lingering in the program. Since I will be on vacation throughout July, I encourage everyone to submit bug reports and questions, but I will not promise to respond to them until in August.
I hope that you will enjoy this new ITSx release, which you can download here. Happy barcoding!
Published paper: Investigating resistomes using metagenomics
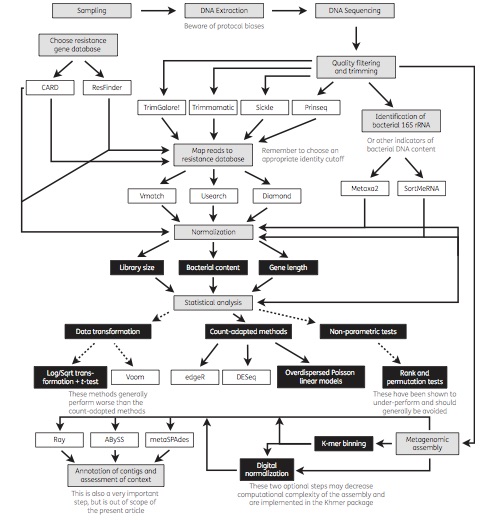
Today, a review paper which I wrote together with Joakim Larsson and Erik Kristiansson was published in Journal of Antimicrobial Chemotherapy (1). We have for a long time used metagenomic DNA sequencing to study antibiotic resistance in different environments (2-6), including in the human microbiota (7). Generally, our ultimate purpose has been to assess the risks to human health associated with resistance genes in the environment. However, a multitude of methods exist for metagenomic data analysis, and over the years we have learned that not all methods are suitable for the investigation of resistance genes for this purpose. In our review paper, we describe and discuss current methods for sequence handling, mapping to databases of resistance genes, statistical analysis and metagenomic assembly. We also provide an overview of important considerations related to the analysis of resistance genes, and end by recommending some of the currently used tools, databases and methods that are best equipped to inform research and clinical practice related to antibiotic resistance (see the figure from the paper below). We hope that the paper will be useful to researchers and clinicians interested in using metagenomic sequencing to better understand the resistance genes present in environmental and human-associated microbial communities.
References
- Bengtsson-Palme J, Larsson DGJ, Kristiansson E: Using metagenomics to investigate human and environmental resistomes. Journal of Antimicrobial Chemotherapy, advance access (2017). doi: 10.1093/jac/dkx199 [Paper link]
- Bengtsson-Palme J, Boulund F, Fick J, Kristiansson E, Larsson DGJ: Shotgun metagenomics reveals a wide array of antibiotic resistance genes and mobile elements in a polluted lake in India. Frontiers in Microbiology, 5, 648 (2014). doi: 10.3389/fmicb.2014.00648 [Paper link]
- Lundström S, Östman M, Bengtsson-Palme J, Rutgersson C, Thoudal M, Sircar T, Blanck H, Eriksson KM, Tysklind M, Flach C-F, Larsson DGJ: Minimal selective concentrations of tetracycline in complex aquatic bacterial biofilms. Science of the Total Environment, 553, 587–595 (2016). doi: 10.1016/j.scitotenv.2016.02.103 [Paper link]
- Bengtsson-Palme J, Hammarén R, Pal C, Östman M, Björlenius B, Flach C-F, Kristiansson E, Fick J, Tysklind M, Larsson DGJ: Elucidating selection processes for antibiotic resistance in sewage treatment plants using metagenomics. Science of the Total Environment, 572, 697–712 (2016). doi: 10.1016/j.scitotenv.2016.06.228 [Paper link]
- Pal C, Bengtsson-Palme J, Kristiansson E, Larsson DGJ: The structure and diversity of human, animal and environmental resistomes. Microbiome, 4, 54 (2016). doi: 10.1186/s40168-016-0199-5 [Paper link]
- Flach C-F, Pal C, Svensson CJ, Kristiansson E, Östman M, Bengtsson-Palme J, Tysklind M, Larsson DGJ: Does antifouling paint select for antibiotic resistance? Science of the Total Environment, 590–591, 461–468 (2017). doi: 10.1016/j.scitotenv.2017.01.213 [Paper link]
- Bengtsson-Palme J, Angelin M, Huss M, Kjellqvist S, Kristiansson E, Palmgren H, Larsson DGJ, Johansson A: The human gut microbiome as a transporter of antibiotic resistance genes between continents. Antimicrobial Agents and Chemotherapy, 59, 10, 6551–6560 (2015). doi: 10.1128/AAC.00933-15 [Paper link]
Published paper: Evaluating taxonomic classification software
Yesterday, Molecular Ecology Resources put online an unedited version of a recent paper which I co-authored. This time, Rodney Richardson at Ohio State University has made a tremendous work of evaluating three taxonomic classification software – the RDP Naïve Bayesian Classifier, RTAX and UTAX – on a set of DNA barcoding regions commonly used for plants, namely the ITS2, and the matK, rbcL, trnL and trnH genes.
In the paper (1), we discuss the results, merits and limitations of the classifiers. In brief, we found that:
- There is a considerable trade-off between accuracy and sensitivity for the classifiers tested, which indicates a need for improved sequence classification tools (2)
- UTAX was superior with respect to error rate, but was exceedingly stringent and thus suffered from a low assignment rate
- The RDP Naïve Bayesian Classifier displayed high sensitivity and low error at the family and order levels, but had a genus-level error rate of 9.6 percent
- RTAX showed high sensitivity at all taxonomic ranks, but at the same time consistently produced the high error rates
- The choice of locus has significant effects on the classification sensitivity of all tested tools
- All classifiers showed strong relationships between database completeness, classification sensitivity and classification accuracy
We believe that the methods of comparison we have used are simple and robust, and thereby provides a methodological and conceptual foundation for future software evaluations. On a personal note, I will thoroughly enjoy working with Rodney and Reed again; I had a great time discussing the ins and outs of taxonomic classification with them! The paper can be found here.
References and notes
- Richardson RT, Bengtsson-Palme J, Johnson RM: Evaluating and Optimizing the Performance of Software Commonly Used for the Taxonomic Classification of DNA Sequence Data. Molecular Ecology Resources, Early view (2016). doi: 10.1111/1755-0998.12628 [Paper link]
- This is something that several classifiers also showed in the evaluation we did for the Metaxa2 paper (3). Interestingly enough, Metaxa2 is better at maintaining high accuracy also as sensitivity is increased.
- Bengtsson-Palme J, Hartmann M, Eriksson KM, Pal C, Thorell K, Larsson DGJ, Nilsson RH: Metaxa2: Improved identification and taxonomic classification of small and large subunit rRNA in metagenomic data. Molecular Ecology Resources, 15, 6, 1403–1414 (2015). doi: 10.1111/1755-0998.12399 [Paper link]
SBW2016
Today the 15th Swedish Bioinformatics Workshop starts in Linköping. For the first time in many many years, I will not be there, which feels super-strange. (I even took part in organizing the workshop two years ago, in Gothenburg.) I would have thoroughly enjoyed the schedule (the workshops look amazing), and hope that everyone will have a very good time there without me. I do miss you! Please make sure I get another opportunity next year!