Blog post at Nature Microbiology Community
I have just published a popular-science-for-scientists type of post at the Nature Microbiology Community about my recent paper published in Microbiome. I personally think that it might be worth a read, so feel free to head over here and read it!
Published paper: Predicting the uncharacterized resistome
Over the weekend, Microbiome put online my most recent paper (1) – a project which started as an idea I got when I finished up my PhD thesis in 2016. One of my main points in the thesis (2), which was also made again on our recent review on environmental factors influencing resistance development (3), is that the greatest risks associated with antibiotic resistance in the environment may not be the resistance genes already circulating in pathogens (which are relatively easily quantified), but the ones associated with recruitment of novel resistance genes from bacteria in the environment (2-4). The latter genes are, however, impossible to quantify due to the fact that they are unknown. But what if we could use knowledge of the diversity and abundance of known resistance genes to estimate the same properties of the yet uncharacterized resistome? That would be a great advantage in e.g. ranking of risk environments, as then some property that is easily monitored can be used to inform risk management of both known and unknown resistance factors.
This just published paper explores this possibility, by quantifying the abundance and diversity of resistance genes in 1109 metagenomes from various environments (1). I have taken two different approaches. First, I took out smaller subsets of genes from the reference database (in this case Resqu, a database of antibiotic resistance genes with verified resistance functions, detected on mobile genetic elements), and used those subsets to estimate resistome diversity and abundance in the 1109 metagenomes. Then these predictions were compared to the results of the entire database. I then, in a second step, investigated if these predictions could be extended to a set of truly novel resistance genes, i.e. the resistance genes present in the FARME database, collecting data from functional metagenomics inserts (5,6).
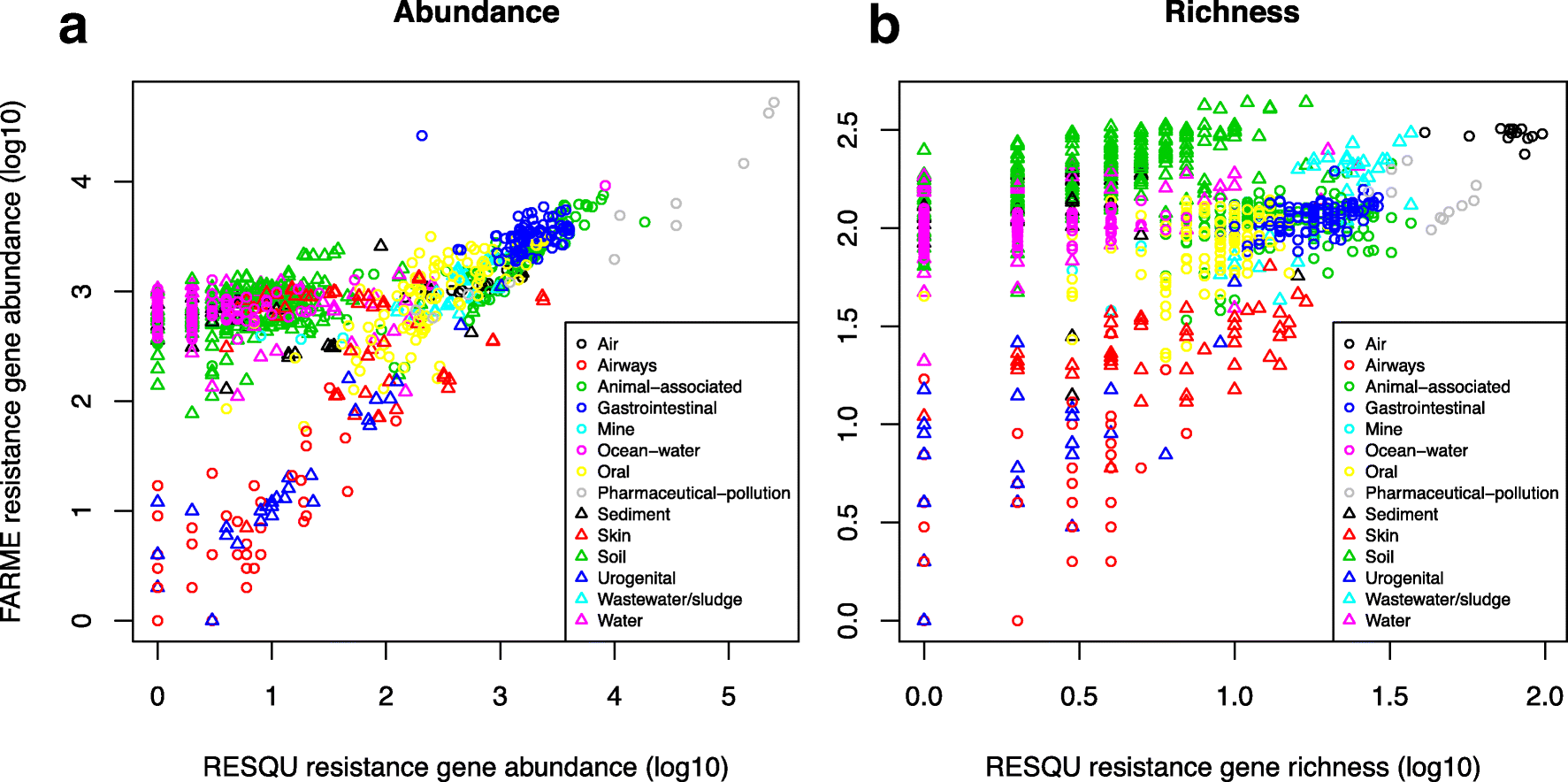
The results show that generally the diversity and abundance of known antibiotic resistance genes can be used to predict the same properties of undescribed resistance genes (see figure above). However, the extent of this predictability is, importantly, dependent on the type of environment investigated. The study also shows that carefully selected small sets of resistance genes can describe total resistance gene diversity remarkably well. This means that knowledge gained from large-scale quantifications of known resistance genes can be utilized as a proxy for unknown resistance factors. This is important for current and proposed monitoring efforts for environmental antibiotic resistance (7-11) and has implications for the design of risk ranking strategies and the choices of measures and methods for describing resistance gene abundance and diversity in the environment.
The study also investigated which diversity measures were best suited to estimate total diversity. Surprisingly, some diversity measures described the total diversity of resistance genes remarkably bad. Most prominently, the Simpson diversity index consistently showed poor performance, and while the Shannon index performed relatively better, there is still no reason to select the Shannon index over normalized (rarefied) richness of resistance genes. The ACE estimator fluctuated substantially compared to the other diversity measures, while the Chao1 estimator more consistently showed performance very similar to richness. Therefore, either richness or the Chao1 estimator should be used for ranking resistance gene diversity, while the Shannon, Simpson, and ACE measures should be avoided.
Importantly, this study implies that the recruitment of novel antibiotic resistance genes from the environment to human pathogens is essentially random. Therefore, when ranking risks associated with antibiotic resistance in environmental settings, the knowledge gained from large-scale quantification of known resistance genes can be utilized as a proxy for the unknown resistance factors (although this proxy is not perfect). Thus, high-risk environments for resistance development and dissemination would, for example, be aquaculture, animal husbandry, discharges from antibiotic manufacturing, and untreated sewage (3,8,12-15). Further attention should probably be paid to antibiotic contaminated soils, as this study points to soils as a vast source of resistance genes not yet encountered in human pathogens. This has also been suggested previously by others (16-19). The results of this study can be used to guide monitoring efforts for environmental antibiotic resistance, to design risk ranking strategies, and to choose appropriate measures and methods for describing resistance gene abundance and diversity in the environment. The entire open access paper is available here.
References
- Bengtsson-Palme J: The diversity of uncharacterized antibiotic resistance genes can be predicted from known gene variants – but not always. Microbiome, 6, 125 (2018). doi: 10.1186/s40168-018-0508-2
- Bengtsson-Palme J: Antibiotic resistance in the environment: a contribution from metagenomic studies. Doctoral thesis (medicine), Department of Infectious Diseases, Institute of Biomedicine, Sahlgrenska Academy, University of Gothenburg, 2016. [Link]
- Bengtsson-Palme J, Kristiansson E, Larsson DGJ: Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, 42, 1, 68–80 (2018). doi: 10.1093/femsre/fux053
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Wallace JC, Port JA, Smith MN, Faustian EM: FARME DB: a functional antibiotic resistance element database. Database, 2017, baw165 (2017).
- Handelsman J, Rondon MR, Brady SF, Clardy J, Goodman RM: Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products. Chemical Biology, 5, R245–249 (1998).
- Berendonk TU, Manaia CM, Merlin C, Fatta-Kassinos D, Cytryn E, Walsh F, et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Pruden A, Larsson DGJ, Amézquita A, Collignon P, Brandt KK, Graham DW, et al.: Management options for reducing the release of antibiotics and antibiotic resistance genes to the environment. Environmental Health Perspectives, 121, 878–885 (2013).
- Review on Antimicrobial Resistance: Antimicrobials in agriculture and the environment: reducing unnecessary use and waste. O’Neill J, ed. London: Wellcome Trust & HM Government (2015).
- Angers-Loustau A, Petrillo M, Bengtsson-Palme J, Berendonk T, Blais B, Chan KG, Coque TM, Hammer P, Heß S, Kagkli DM, Krumbiegel C, Lanza VF, Madec J-Y, Naas T, O’Grady J, Paracchini V, Rossen JWA, Ruppé E, Vamathevan J, Venturi V, Van den Eede G: The challenges of designing a benchmark strategy for bioinformatics pipelines in the identification of antimicrobial resistance determinants using next generation sequencing technologies. F1000Research, 7, 459 (2018). doi: 10.12688/f1000research.14509.1
- Larsson DGJ, Andremont A, Bengtsson-Palme J, Brandt KK, de Roda Husman AM, Fagerstedt P, Fick J, Flach C-F, Gaze WH, Kuroda M, Kvint K, Laxminarayan R, Manaia CM, Nielsen KM, Ploy M-C, Segovia C, Simonet P, Smalla K, Snape J, Topp E, van Hengel A, Verner-Jeffreys DW, Virta MPJ, Wellington EM, Wernersson A-S: Critical knowledge gaps and research needs related to the environmental dimensions of antibiotic resistance. Environment International, 117, 132–138 (2018). doi: 10.1016/j.envint.2018.04.041
- Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J: Call of the wild: antibiotic resistance genes in natural environments. Nature Reviews Microbiology, 8, 251–259 (2010).
- Graham DW, Collignon P, Davies J, Larsson DGJ, Snape J: Underappreciated role of regionally poor water quality on globally increasing antibiotic resistance. Environmental Science & Technology, 48,11746–11747 (2014).
- Larsson DGJ: Pollution from drug manufacturing: review and perspectives. Philosophical Transactions of the Royal Society of London, Series B Biological Sciences, 369, 20130571 (2014).
- Cabello FC, Godfrey HP, Buschmann AH, Dölz HJ: Aquaculture as yet another environmental gateway to the development and globalisation of antimicrobial resistance. Lancet Infectious Diseases, 16, e127–133 (2016).
- Forsberg KJ, Reyes A, Wang B, Selleck EM, Sommer MOA, Dantas G: The shared antibiotic resistome of soil bacteria and human pathogens. Science, 337, 1107–1111 (2012).
- Allen HK, Moe LA, Rodbumrer J, Gaarder A, Handelsman J: Functional metagenomics reveals diverse beta-lactamases in a remote Alaskan soil. ISME Journal, 3, 243–251 (2009).
- Riesenfeld CS, Goodman RM, Handelsman J: Uncultured soil bacteria are a reservoir of new antibiotic resistance genes. Environmental Microbiology, 6, 981–989 (2004).
- McGarvey KM, Queitsch K, Fields S: Wide variation in antibiotic resistance proteins identified by functional metagenomic screening of a soil DNA library. Applied and Environmental Microbiology, 78, 1708–1714 (2012).
Published paper: Knowledge gaps for environmental antibiotic resistance
The outcomes from a workshop arranged by JPIAMR, the Swedish Research Council (VR) and CARe were just published as a short review paper in Environment International. In the paper, which was mostly moved forward by Prof. Joakim Larsson at CARe, we describe four major areas of knowledge gaps in the realm of environmental antibiotic resistance (1). We then highlight several important sub-questions within these areas. The broad areas we define are:
- What are the relative contributions of different sources of antibiotics and antibiotic resistant bacteria into the environment?
- What is the role of the environment as affected by anthropogenic inputs (e.g. pollution and other activities) on the evolution (mobilization, selection, transfer, persistence etc.) of antibiotic resistance?
- How significant is the exposure of humans to antibiotic resistant bacteria via different environmental routes, and what is the impact on human health?
- What technological, social, economic and behavioral interventions are effective to mitigate the emergence and spread of antibiotic resistance via the environment?
Although much has been written on the topic before (e.g. 2-12), I think it is unique that we collect and explicitly point out areas in which we are lacking important knowledge to build accurate risk models and devise appropriate intervention strategies. The workshop was held in Gothenburg on the 27–28th of September 2017. The workshop leaders Joakim Larsson, Ana-Maria de Roda Husman and Ramanan Laxminarayan were each responsible for moderating a breakout group, and every breakout group was tasked to deal with knowledge gaps related to either evolution, transmission or interventions. The reports of the breakout groups were then discussed among all participants to clarify and structure the areas where more research is needed, which boiled down to the four overarching critical knowledge gaps described in the paper (1).
This is a short paper, and I think everyone with an interest in environmental antibiotic resistance should read it and reflect over its content (because, we may of course have overlooked some important aspect). You can find the paper here.
References
- Larsson DGJ, Andremont A, Bengtsson-Palme J, Brandt KK, de Roda Husman AM, Fagerstedt P, Fick J, Flach C-F, Gaze WH, Kuroda M, Kvint K, Laxminarayan R, Manaia CM, Nielsen KM, Ploy M-C, Segovia C, Simonet P, Smalla K, Snape J, Topp E, van Hengel A, Verner-Jeffreys DW, Virta MPJ, Wellington EM, Wernersson A-S: Critical knowledge gaps and research needs related to the environmental dimensions of antibiotic resistance. Environment International, 117, 132–138 (2018). doi: 10.1016/j.envint.2018.04.041
- Bengtsson-Palme J, Kristiansson E, Larsson DGJ: Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, 42, 1, 68–80 (2018). doi: 10.1093/femsre/fux053
- Martinez JL, Coque TM, Baquero F: What is a resistance gene? Ranking risk in resistomes. Nature Reviews Microbiology 2015, 13:116–123. doi:10.1038/nrmicro3399
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Ashbolt NJ, Amézquita A, Backhaus T, Borriello P, Brandt KK, Collignon P, et al.: Human Health Risk Assessment (HHRA) for Environmental Development and Transfer of Antibiotic Resistance. Environmental Health Perspectives, 121, 993–1001 (2013)
- Pruden A, Larsson DGJ, Amézquita A, Collignon P, Brandt KK, Graham DW, et al.: Management options for reducing the release of antibiotics and antibiotic resistance genes to the environment. Environmental Health Perspectives, 121, 878–85 (2013).
- Gillings MR: Evolutionary consequences of antibiotic use for the resistome, mobilome and microbial pangenome. Frontiers in Microbiology, 4, 4 (2013).
- Baquero F, Alvarez-Ortega C, Martinez JL: Ecology and evolution of antibiotic resistance. Environmental Microbiology Reports, 1, 469–476 (2009).
- Baquero F, Tedim AP, Coque TM: Antibiotic resistance shaping multi-level population biology of bacteria. Frontiers in Microbiology, 4, 15 (2013).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Hiltunen T, Virta M, Laine A-L: Antibiotic resistance in the wild: an eco-evolutionary perspective. Philosophical Transactions of the Royal Society B: Biological Sciences, 372 (2017) doi: 10.1098/rstb.2016.0039.
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
New preprint: benchmarking resistance gene identification
This weekend, F1000Research put online the non-peer-reviewed version of the paper resulting from a workshop arranged by the JRC in Italy last year (1). (I will refer to this as a preprint, but at F1000Research the line is quite blurry between preprint and published paper.) The paper describes various challenges arising from the process of designing a benchmark strategy for bioinformatics pipelines (2) in the identification of antimicrobial resistance genes in next generation sequencing data.
The paper discusses issues about the benchmarking datasets used, testing samples, evaluation criteria for the performance of different tools, and how the benchmarking dataset should be created and distributed. Specially, we address the following questions:
- How should a benchmark strategy handle the current and expanding universe of NGS platforms?
- What should be the quality profile (in terms of read length, error rate, etc.) of in silico reference materials?
- Should different sets of reference materials be produced for each platform? In that case, how to ensure no bias is introduced in the process?
- Should in silico reference material be composed of the output of real experiments, or simulated read sets? If a combination is used, what is the optimal ratio?
- How is it possible to ensure that the simulated output has been simulated “correctly”?
- For real experiment datasets, how to avoid the presence of sensitive information?
- Regarding the quality metrics in the benchmark datasets (e.g. error rate, read quality), should these values be fixed for all datasets, or fall within specific ranges? How wide can/should these ranges be?
- How should the benchmark manage the different mechanisms by which bacteria acquire resistance?
- What is the set of resistance genes/mechanisms that need to be included in the benchmark? How should this set be agreed upon?
- Should datasets representing different sample types (e.g. isolated clones, environmental samples) be included in the same benchmark?
- Is a correct representation of different bacterial species (host genomes) important?
- How can the “true” value of the samples, against which the pipelines will be evaluated, be guaranteed?
- What is needed to demonstrate that the original sample has been correctly characterised, in case real experiments are used?
- How should the target performance thresholds (e.g. specificity, sensitivity, accuracy) for the benchmark suite be set?
- What is the impact of these performance thresholds on the required size of the sample set?
- How can the benchmark stay relevant when new resistance mechanisms are regularly characterized?
- How is the continued quality of the benchmark dataset ensured?
- Who should generate the benchmark resource?
- How can the benchmark resource be efficiently shared?
Of course, we have not answered all these questions, but I think we have come down to a decent description of the problems, which we see as an important foundation for solving these issues and implementing the benchmarking standard. Some of these issues were tackled in our review paper from last year on using metagenomics to study resistance genes in microbial communities (3). The paper also somewhat connects to the database curation paper we published in 2016 (4), although this time the strategies deal with the testing datasets rather than the actual databases. The paper is the first outcome of the workshop arranged by the JRC on “Next-generation sequencing technologies and antimicrobial resistance” held October 4-5 last year in Ispra, Italy. You can find the paper here (it’s open access).
References and notes
- Angers-Loustau A, Petrillo M, Bengtsson-Palme J, Berendonk T, Blais B, Chan KG, Coque TM, Hammer P, Heß S, Kagkli DM, Krumbiegel C, Lanza VF, Madec J-Y, Naas T, O’Grady J, Paracchini V, Rossen JWA, Ruppé E, Vamathevan J, Venturi V, Van den Eede G: The challenges of designing a benchmark strategy for bioinformatics pipelines in the identification of antimicrobial resistance determinants using next generation sequencing technologies. F1000Research, 7, 459 (2018). doi: 10.12688/f1000research.14509.1
- You may remember that I hate the term “pipeline” for bioinformatics protocols. I would have preferred if it was called workflows or similar, but the term “pipeline” has taken hold and I guess this is a battle where I have essentially lost. The bioinformatics workflows will be known as pipelines, for better and worse.
- Bengtsson-Palme J, Larsson DGJ, Kristiansson E: Using metagenomics to investigate human and environmental resistomes. Journal of Antimicrobial Chemotherapy, 72, 2690–2703 (2017). doi: 10.1093/jac/dkx199
- Bengtsson-Palme J, Boulund F, Edström R, Feizi A, Johnning A, Jonsson VA, Karlsson FH, Pal C, Pereira MB, Rehammar A, Sánchez J, Sanli K, Thorell K: Strategies to improve usability and preserve accuracy in biological sequence databases. Proteomics, 16, 18, 2454–2460 (2016). doi: 10.1002/pmic.201600034
An advice to journal editors
I was recently invited to review a manuscript for a journal (1). After half the time to review deadline had passed, I received a mail stating that “In the interest of your time and the authors’ time, I am making a decision without the benefit of your input.” While I do understand that big journals receive many submissions and that the editor made this decision in the interest of time, I think that it should also be kept in mind that I had already spent approximately three hours scrutinizing the manuscript. Due to the decision of the editor, these are now three hours of work down the drain.
Furthermore, I was not informed what the decision was, and my access to the paper in the reviewing system was revoked. This means that I don’t even know if my opinions are concordant with the rest of the reviewers or not. Perhaps I had actually spotted crucial errors that the other reviewers had missed? Or maybe the paper was rejected, and my input was therefore no longer needed. I don’t know, because I was not informed.
These days, I receive many requests to perform manuscript reviews. A journal treating its reviewers like this causes me to lose all willingness to review for that journal again. To me, making a decision to dismiss reviewers without even asking them if they are about to submit comments, signals that a journal does not value its peer reviewers, and that I can spend my time better elsewhere. Similar to authors and editors, I do not want to waste my time on tasks that end up being of no use.
On the upside, this decision by the editor has freed some time for me to write up this rant, including the following advice: If you are an editor of a journal and you want to keep the reviewers (who, I remind you, work for free and are largely unrecognized for their work) happy, try to avoid pissing them off by dismissing their work. It does not hurt to ask them if they are about to submit their comments, or if they – given that a sufficient number of review reports have been submitted – would rather withdraw from the review process. This may add an extra day or two to the process, but I think that in the long run both authors, editors and reviewers would agree that the overall quality of the peer review process would benefit from those few extra days.
Footnotes
- I am not going to name the journal here, nor the identity of the editor, because that is not my point. I am not after singling out certain people here, but I want to address an overall behavior that annoys me. That said, the last three papers I have been a co-author on in this journal took five to seven weeks from the authors correction of the proofs until publication online. I find it stunning that with these delays, the journal dismisses the reviewers it has invited because they don’t produce peer reviews quicker than the deadline proposed by the journal. The real bottleneck in this process is not extensive review times, at least not in my experience.
The Wisconsin Blog is on again
I’ve been having a very intense start of the year with the move to the US and getting the family accustomed to Madison (which has taken time and energy, but gone really well). I just wanted to make you aware of that I have started posting at the Wisconsin Blog again and hope to be sharing research related stuff from my year in the US there. For more personal stuff, our family has set up a blog (in Swedish) at this address: https://palmeiamerikat.blogspot.com. You are very welcome to follow our adventure there!
Published paper: Annotating fungi from the built environment part II
MycoKeys earlier this week published a paper describing the results of a workshop in Aberdeen in April last year, where we refined annotations for fungal ITS sequences from the built environment (1). This was a follow-up on a workshop in May 2016 (2) and the results have been implemented in the UNITE database and shared with other online resources. The paper has also been highlighted at microBEnet. I have very little time to further comment on this at this very moment, but I believe, as I wrote last time, that distributed initiatives like this (and the ones I have been involved in in the past (3,4)) serve a very important purpose for establishing better annotation of sequence data (5). The full paper can be found here.
References
- Nilsson RH, Taylor AFS, Adams RI, Baschien C, Bengtsson-Palme J, Cangren P, Coleine C, Daniel H-M, Glassman SI, Hirooka Y, Irinyi L, Iršenaite R, Martin-Sánchez PM, Meyer W, Oh S-O, Sampaio JP, Seifert KA, Sklenár F, Stubbe D, Suh S-O, Summerbell R, Svantesson S, Unterseher M, Visagie CM, Weiss M, Woudenberg J, Wurzbacher C, Van den Wyngaert S, Yilmaz N, Yurkov A, Kõljalg U, Abarenkov K: Annotating public fungal ITS sequences from the built environment according to the MIxS-Built Environment standard – a report from an April 10-11, 2017 workshop (Aberdeen, UK). MycoKeys, 28, 65–82 (2018). doi: 10.3897/mycokeys.28.20887 [Paper link]
- Abarenkov K, Adams RI, Laszlo I, Agan A, Ambrioso E, Antonelli A, Bahram M, Bengtsson-Palme J, Bok G, Cangren P, Coimbra V, Coleine C, Gustafsson C, He J, Hofmann T, Kristiansson E, Larsson E, Larsson T, Liu Y, Martinsson S, Meyer W, Panova M, Pombubpa N, Ritter C, Ryberg M, Svantesson S, Scharn R, Svensson O, Töpel M, Untersehrer M, Visagie C, Wurzbacher C, Taylor AFS, Kõljalg U, Schriml L, Nilsson RH: Annotating public fungal ITS sequences from the built environment according to the MIxS-Built Environment standard – a report from a May 23-24, 2016 workshop (Gothenburg, Sweden). MycoKeys, 16, 1–15 (2016). doi: 10.3897/mycokeys.16.10000
- Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M, Bates ST, Bruns TT, Bengtsson-Palme J, Callaghan TM, Douglas B, Drenkhan T, Eberhardt U, Dueñas M, Grebenc T, Griffith GW, Hartmann M, Kirk PM, Kohout P, Larsson E, Lindahl BD, Lücking R, Martín MP, Matheny PB, Nguyen NH, Niskanen T, Oja J, Peay KG, Peintner U, Peterson M, Põldmaa K, Saag L, Saar I, Schüßler A, Senés C, Smith ME, Suija A, Taylor DE, Telleria MT, Weiß M, Larsson KH: Towards a unified paradigm for sequence-based identification of Fungi. Molecular Ecology, 22, 21, 5271–5277 (2013). doi: 10.1111/mec.12481
- Nilsson RH, Hyde KD, Pawlowska J, Ryberg M, Tedersoo L, Aas AB, Alias SA, Alves A, Anderson CL, Antonelli A, Arnold AE, Bahnmann B, Bahram M, Bengtsson-Palme J, Berlin A, Branco S, Chomnunti P, Dissanayake A, Drenkhan R, Friberg H, Frøslev TG, Halwachs B, Hartmann M, Henricot B, Jayawardena R, Jumpponen A, Kauserud H, Koskela S, Kulik T, Liimatainen K, Lindahl B, Lindner D, Liu J-K, Maharachchikumbura S, Manamgoda D, Martinsson S, Neves MA, Niskanen T, Nylinder S, Pereira OL, Pinho DB, Porter TM, Queloz V, Riit T, Sanchez-García M, de Sousa F, Stefaczyk E, Tadych M, Takamatsu S, Tian Q, Udayanga D, Unterseher M, Wang Z, Wikee S, Yan J, Larsson E, Larsson K-H, Kõljalg U, Abarenkov K: Improving ITS sequence data for identification of plant pathogenic fungi. Fungal Diversity, 67, 1, 11–19 (2014). doi: 10.1007/s13225-014-0291-8
- Bengtsson-Palme J, Boulund F, Edström R, Feizi A, Johnning A, Jonsson VA, Karlsson FH, Pal C, Pereira MB, Rehammar A, Sánchez J, Sanli K, Thorell K: Strategies to improve usability and preserve accuracy in biological sequence databases. Proteomics, Early view (2016). doi: 10.1002/pmic.201600034
New beta brings major Metaxa2 updates
I am very happy to announce that a first public beta version of Metaxa2 version 2.2 has been released today! This new version brings two big and a number of small improvements to the Metaxa2 software (1). The first major addition is the introduction of the Metaxa2 Database Builder, which allows the user to create custom databases for virtually any genetic barcoding region. The second addition, which is related to the first, is that the classifier has been rewritten to have a more solid mathematical foundation. I have been promising that these updates were coming “soon” for one and a half years, but finally the end-product is good enough to see some real world testing. Bear in mind though that this is still a beta version that could contain obscure bugs. Here follows a list of new features (with further elaboration on a few below):
- The Metaxa2 Database Builder
- Support for additional barcoding genes, virtually any genetic region can now be used for taxonomic classification in Metaxa2
- The Metaxa2 database repository, which can be accessed through the new metaxa2_install_database tool
- Improved classification scoring model for better clarity and sensitivity
- A bundled COI database for athropods, showing off the capabilities of the database builder
- Support for compressed input files (gzip, zip, bzip, dsrc)
- Support for auto-detection of database locations
- Added output of probable taxonomic origin for sequences with reliability scores at each rank, made possible by the updated classifier
- Added the -x option for running only the extraction without the classification step
- Improved memory handling for very large rRNA datasets in the classifier (millions of sequences)
- This update also fixes a bug in the metaxa2_rf tool that could cause bias in very skewed datasets with small numbers of taxa
The new version of Metaxa2 can be downloaded here, and for those interested I will spend the rest of this post outlining the Metaxa2 Database Builder. The information below is also available in a slightly extended version in the software manual.
The major enhancement in Metaxa2 version 2.2 is the ability to use custom databases for classification. This means that the user can now make their own database for their own barcoding region of choice, or download additional databases from the Metaxa2 Database Repository. The selection of other databases is made through the “-g” option already existing in Metaxa2. As part of these changes, we have also updated the classification scoring model for better stringency and sensitivity across multiple databases and different genes. The old scoring system can still be used by specifying the –scoring_model option to “old”.
There are two different main operating modes of the Metaxa2 Database Builder, as well as a hybrid mode combining the features of the two other modes. The divergent and conserved modes work in almost completely different ways and deal with two different types of barcoding regions. The divergent mode is designed to deal with barcoding regions that exhibit fairly large variation between taxa within the same taxonomic domain. Such regions include, e.g., the eukaryotic ITS region, or the trnL gene used for plant barcoding. In the other mode – the conserved mode – a highly conserved barcoding region is expected (at least within the different taxonomic domains). Genes that fall into this category would be, e.g., the 16S SSU rRNA, and the bacterial rpoB gene. This option would most likely also be suitable for barcoding within certain groups of e.g. plants, where similarity of the barcoding regions can be expected to be high. There is also a third mode – the hybrid mode – that incorporates features of both the other. The hybrid mode is more experimental in nature, but could be useful in situations where both the other modes perform poorer than desired.
In the divergent (default) mode, the database builder will start by clustering the input sequences at 20% identity using USEARCH (2). All clusters generated from this process are then individually aligned using MAFFT (3). Those alignments are split into two regions, which are used to build two hidden Markov models for each cluster of sequences. These models will be less precise, but more sensitive than those generated in the conserved mode. In the divergent mode, the database builder will attempt to extract full-length sequences from the input data, but this may be less successful than in the conserved mode.
In the conserved mode, on the other hand, the database builder will first extract the barcoding region from the input sequences using models built from a reference sequence provided (see above) and the Metaxa2 extractor (1). It will then align all the extracted sequences using MAFFT and determine the conservation of each position in the alignment. When the criteria for degree of conservation are met, all conserved regions are extracted individually and are then re-aligned separately using MAFFT. The re-aligned sequences are used to build hidden Markov models representing the conserved regions with HMMER (4). In this mode, the classification database will only consist of the extracted full-length sequences.
In the hybrid mode, finally, the database builder will cluster the input sequences at 20% identity using USEARCH, and then proceed with the conserved mode approach on each cluster separately .
The actual taxonomic classification in Metaxa2 is done using a sequence database. It was shown in the original Metaxa2 paper that replacing the built-in database with a generic non-processed one was detrimental to performance in terms of accuracy (1). In the database builder, we have tried to incorporate some of the aspects of the manual database curation we did for the built-in database that can be automated. By default, all these filtration steps are turned off, but enabling them might drastically increase the accuracy of classifications based on the database.
To assess the accuracy of the constructed database, the Metaxa2 Database Builder allows for testing the detection ability and classification accuracy of the constructed database. This is done by sub-dividing the database sequences into subsets and rebuilding the database using a smaller (by default 90%), randomly selected, set of the sequence data (5). The remaining sequences (10% by default) are then classified using Metaxa2 with the subset database. The number of detections, and the numbers of correctly or incorrectly classified entries are recorded and averaged over a number of iterations (10 by default). This allows for obtaining a picture of the lower end of the accuracy of the database. However, since the evaluation only uses a subset of all sequences included in the full database, the performance of the full database actually constructed is likely to be slightly better. The evaluation can be turned on using the “–evaluate T” option.
Metaxa2 2.2 also introduces the database repository, from which the user can download additional databases for Metaxa2. To download new databases from the repository, the metaxa2_install_database command is used. This is a simple piece of software but requires internet access to function.
References
- Bengtsson-Palme J, Hartmann M, Eriksson KM, Pal C, Thorell K, Larsson DGJ, Nilsson RH: Metaxa2: Improved Identification and Taxonomic Classification of Small and Large Subunit rRNA in Metagenomic Data. Molecular Ecology Resources (2015). doi: 10.1111/1755-0998.12399 [Paper link]
- Edgar RC: Search and clustering orders of magnitude faster than BLAST. Bioinformatics, 26, 2460–2461 (2010).
- Katoh K, Standley DM: MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution, 30, 772–780 (2013).
- Eddy SR: Accelerated profile HMM searches. PLoS Computational Biology, 7, e1002195 (2011).
- Richardson RT, Bengtsson-Palme J, Johnson RM: Evaluating and Optimizing the Performance of Software Commonly Used for the Taxonomic Classification of DNA Sequence Data. Molecular Ecology Resources, 17, 4, 760–769 (2017). doi: 10.1111/1755-0998.12628
Published paper: Environmental factors leading to resistance
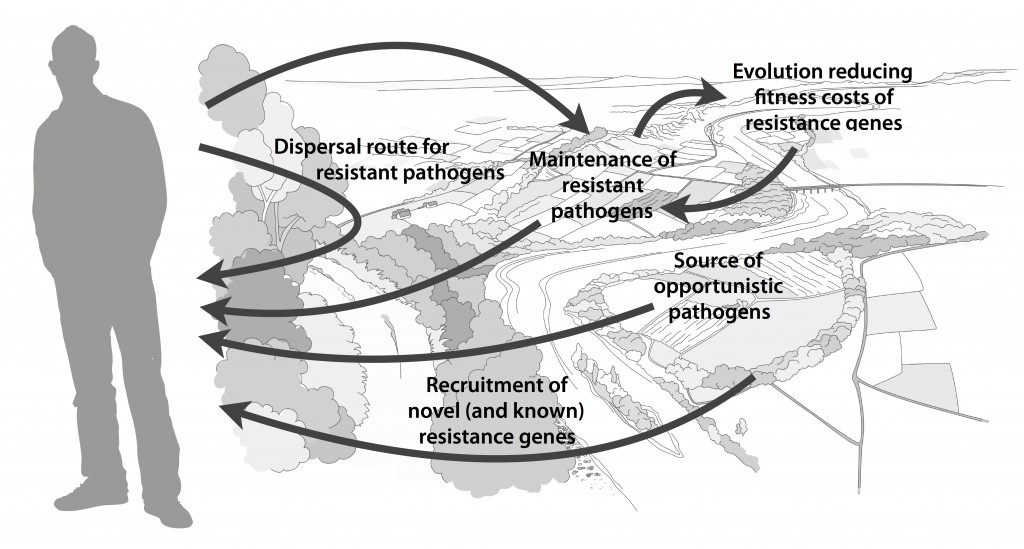
Myself, Joakim Larsson and Erik Kristiansson have written a review on the environmental factors that influence development and spread of antibiotic resistance, which was published today in FEMS Microbiology Reviews. The review (1) builds on thoughts developed in the latter parts of my PhD thesis (2), and seeks to provide a synthesis knowledge gained from different subfields towards the current understanding of evolutionary and ecological processes leading to clinical appearance of resistance genes, as well as the important environmental dispersal barriers preventing spread of resistant pathogens.
We postulate that emergence of novel resistance factors and mobilization of resistance genes are likely to occur continuously in the environment. However, the great majority of such genetic events are unlikely to lead to establishment of novel resistance factors in bacterial populations, unless there is a selection pressure for maintaining them or their fitness costs are negligible. To enable measures to prevent resistance development in the environment, it is therefore critical to investigate under what conditions and to what extent environmental selection for resistance takes place. Selection for resistance is likely less important for the dissemination of resistant bacteria, but will ultimately depend on how well the species or strain in question thrives in the external environment. Metacommunity theory (3,4) suggests that dispersal ability is central to this process, and therefore opportunistic pathogens with their main habitat in the environment may play an important role in the exchange of resistance factors between humans and the environment. Understanding the dispersal barriers hindering this exchange is not only key to evaluate risks, but also to prevent resistant pathogens, as well as novel resistance genes, from reaching humans.
Towards the end of the paper, we suggest certain environments that seem to be more important from a risk management perspective. We also discuss additional problems linked to the development of antibiotic resistance, such as increased evolvability of bacterial genomes (5) and which other types of genes that may be mobilized in the future, should the development continue (1,6). In this review, we also further develop thoughts on the relative risks of re-recruiting and spreading well-known resistance factors already circulating in pathogens, versus recruitment of completely novel resistance genes from environmental bacteria (7). While the latter case is likely to be very rare, and thus almost impossible to quantify the risks for, the consequences of such (potentially one-time) events can be dire.
I personally think that this is one of the best though-through pieces I have ever written, and since it is open access and (in my biased opinion) written in a fairly accessible way, I recommend everyone to read it. It builds on the ecological theories for resistance ecology developed by, among others, Fernando Baquero and José Martinez (8-13). Over the last year, it has been stressed several times at meetings (e.g. at the EDAR conferences in August) that there is a need to develop an ecological framework for antibiotic resistance genes. I think this paper could be one of the foundational pillars on such an endeavor and look forward to see how it will fit into the growing literature on the subject!
References
- Bengtsson-Palme J, Kristiansson E, Larsson DGJ: Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, accepted manuscript (2017). doi: 10.1093/femsre/fux053
- Bengtsson-Palme J: Antibiotic resistance in the environment: a contribution from metagenomic studies. Doctoral thesis (medicine), Department of Infectious Diseases, Institute of Biomedicine, Sahlgrenska Academy, University of Gothenburg, 2016. [Link]
- Bengtsson J: Applied (meta)community ecology: diversity and ecosystem services at the intersection of local and regional processes. In: Verhoef HA, Morin PJ (eds.). Community Ecology: Processes, Models, and Applications. Oxford: Oxford University Press, 115–130 (2009).
- Leibold M, Norberg J: Biodiversity in metacommunities: Plankton as complex adaptive systems? Limnology and Oceanography, 1278–1289 (2004).
- Gillings MR, Stokes HW: Are humans increasing bacterial evolvability? Trends in Ecology and Evolution, 27, 346–352 (2012).
- Gillings MR: Evolutionary consequences of antibiotic use for the resistome, mobilome and microbial pangenome. Frontiers in Microbiology, 4, 4 (2013).
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Baquero F, Alvarez-Ortega C, Martinez JL: Ecology and evolution of antibiotic resistance. Environmental Microbiology Reports, 1, 469–476 (2009).
- Baquero F, Tedim AP, Coque TM: Antibiotic resistance shaping multi-level population biology of bacteria. Frontiers in Microbiology, 4, 15 (2013).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Hiltunen T, Virta M, Laine A-L: Antibiotic resistance in the wild: an eco-evolutionary perspective. Philosophical Transactions of the Royal Society B: Biological Sciences, 372 (2017) doi: 10.1098/rstb.2016.0039.
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
- Salyers AA, Amábile-Cuevas CF: Why are antibiotic resistance genes so resistant to elimination? Antimicrobial Agents and Chemotherapy, 41, 2321–2325 (1997).
The Wisconsin Blog
I just wanted to notify anyone who might be interested in following my more personal reflections on my month in Wisconsin (and in Michigan over EDAR4) that I will be updating my Wisconsin Blog at this site (hopefully) regularly. The blog updates are not visible on the first page, so you will have to actively go to the Wisconsin Blog page by clicking in the upper right of the page.