Published paper: The Calanus glacialis mitogenome
Mitochondrial DNA Part B today published a mitochondrial genome announcement paper (1) in which I was involved in doing the assemblies and annotating them. The paper describes the mitogenome of Calanus glacialis, a marine planktonic copepod, which is a keystone species in the Arctic Ocean. The mitogenome is 20,674 bp long, and includes 13 protein-coding genes, 2 rRNA genes and 22 tRNA genes. While this is of course note a huge paper, we believe that this new resource will be of interest in understanding the structure and dynamics of C. glacialis populations. The main work in this paper has been carried out by Marvin Choquet at Nord University in Bodø, Norway. So hats off to him for great work, thanks Marvin! The paper can be read here.
Reference
- Choquet M, Alves Monteiro HJ, Bengtsson-Palme J, Hoarau G: The complete mitochondrial genome of the copepod Calanus glacialis. Mitochondrial DNA Part B, 2, 2, 506–507 (2017). doi: 10.1080/23802359.2017.1361357 [Paper link]
Published paper: Investigating resistomes using metagenomics
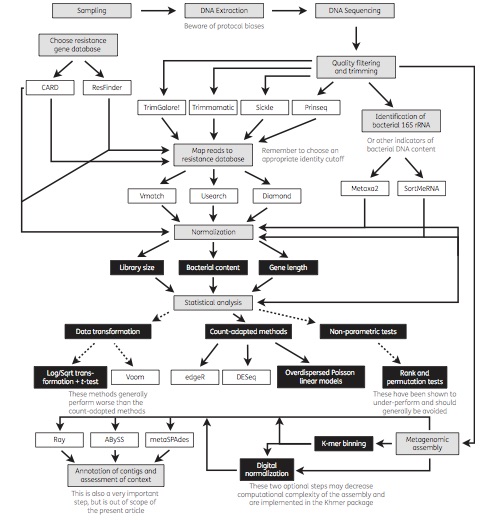
Today, a review paper which I wrote together with Joakim Larsson and Erik Kristiansson was published in Journal of Antimicrobial Chemotherapy (1). We have for a long time used metagenomic DNA sequencing to study antibiotic resistance in different environments (2-6), including in the human microbiota (7). Generally, our ultimate purpose has been to assess the risks to human health associated with resistance genes in the environment. However, a multitude of methods exist for metagenomic data analysis, and over the years we have learned that not all methods are suitable for the investigation of resistance genes for this purpose. In our review paper, we describe and discuss current methods for sequence handling, mapping to databases of resistance genes, statistical analysis and metagenomic assembly. We also provide an overview of important considerations related to the analysis of resistance genes, and end by recommending some of the currently used tools, databases and methods that are best equipped to inform research and clinical practice related to antibiotic resistance (see the figure from the paper below). We hope that the paper will be useful to researchers and clinicians interested in using metagenomic sequencing to better understand the resistance genes present in environmental and human-associated microbial communities.
References
- Bengtsson-Palme J, Larsson DGJ, Kristiansson E: Using metagenomics to investigate human and environmental resistomes. Journal of Antimicrobial Chemotherapy, advance access (2017). doi: 10.1093/jac/dkx199 [Paper link]
- Bengtsson-Palme J, Boulund F, Fick J, Kristiansson E, Larsson DGJ: Shotgun metagenomics reveals a wide array of antibiotic resistance genes and mobile elements in a polluted lake in India. Frontiers in Microbiology, 5, 648 (2014). doi: 10.3389/fmicb.2014.00648 [Paper link]
- Lundström S, Östman M, Bengtsson-Palme J, Rutgersson C, Thoudal M, Sircar T, Blanck H, Eriksson KM, Tysklind M, Flach C-F, Larsson DGJ: Minimal selective concentrations of tetracycline in complex aquatic bacterial biofilms. Science of the Total Environment, 553, 587–595 (2016). doi: 10.1016/j.scitotenv.2016.02.103 [Paper link]
- Bengtsson-Palme J, Hammarén R, Pal C, Östman M, Björlenius B, Flach C-F, Kristiansson E, Fick J, Tysklind M, Larsson DGJ: Elucidating selection processes for antibiotic resistance in sewage treatment plants using metagenomics. Science of the Total Environment, 572, 697–712 (2016). doi: 10.1016/j.scitotenv.2016.06.228 [Paper link]
- Pal C, Bengtsson-Palme J, Kristiansson E, Larsson DGJ: The structure and diversity of human, animal and environmental resistomes. Microbiome, 4, 54 (2016). doi: 10.1186/s40168-016-0199-5 [Paper link]
- Flach C-F, Pal C, Svensson CJ, Kristiansson E, Östman M, Bengtsson-Palme J, Tysklind M, Larsson DGJ: Does antifouling paint select for antibiotic resistance? Science of the Total Environment, 590–591, 461–468 (2017). doi: 10.1016/j.scitotenv.2017.01.213 [Paper link]
- Bengtsson-Palme J, Angelin M, Huss M, Kjellqvist S, Kristiansson E, Palmgren H, Larsson DGJ, Johansson A: The human gut microbiome as a transporter of antibiotic resistance genes between continents. Antimicrobial Agents and Chemotherapy, 59, 10, 6551–6560 (2015). doi: 10.1128/AAC.00933-15 [Paper link]
Report on JRC AMR workshop
In March, I attended a workshop on the role of NGS technologies in the coordinated action plan against antimicrobial resistance, organised by JRC in Italy. I was, together with 14 other experts, invited to discuss where and how sequencing can be used to investigate and manage antibiotic resistance. The report from the workshop has just recently been published, and is available here. There will be follow-up activities on this workshop, which I also hope that I will be able to participate in, since this is an important and very interesting pet topic of mine.
Reference
Webinar: The (un)recognised pathways of AMR
Sorry for the late notice, but if you have half an hour to spare later today I will discuss our findings on resistance genes in Beijing air on a webinar organised by Healthcare Without Harm on “The (un)recognised pathways of AMR: Air pollution and food“. Tune in a few minutes before 16.00 CEST!
Published paper: Does antifouling paint select for antibiotic resistance?
After the usual (1,2) long wait between acceptance and publication, Science of the Total Environment today put a paper online in which I have played a role in the bioinformatic analysis. In the paper, we investigate whether antifouling paint containing copper and zinc could co-select for antibiotic resistance, using microbiological methods and metagenomic sequencing (3).
In this work, we have studied marine microbial biofilms allowed to grow on surfaces painted with antifouling paint submerged in sea water. Such antifouling paints often contain metals that potentially could co-select for antibiotic resistance (4). Using microbiological culturing, we found that the heavy-metal based paint co-selected for bacteria resistant to tetracycline. However, the paint did not enrich neither the total abundance of known mobile antibiotic resistance genes nor the abundance of tetracycline resistance genes in the biofilm communities. Rather, the communities from the painted surfaces were enriched for bacteria with genetic profiles suggesting increased capacity for extrusion of antibiotics via RND efflux systems. In addition, these communities were also enriched for genes involved in mobilization of DNA, such as ISCR transposases and integrases. Finally, the biofilm communities from painted surfaces displayed lower taxonomic diversity and were at the same time enriched for Gammaproteobacteria. The paper builds on our previous work in which we identify certain co-occurences between genes conferring metal and antibiotic resistance (4). However, the findings of this paper do not lend support for that mobile resistance genes are co-selected for by copper and zinc in the marine environment – rather the increase in antibiotic resistance seem to be due to taxonomic changes and cross-resistance mechanisms. The entire paper can be read here.
References
- Bengtsson-Palme J: Published paper: Community MSCs for tetracycline. https://microbiology.se/2016/03/22/published-paper-community-mscs-for-tetracycline/
- Bengtsson-Palme J: Published paper: Antibiotic resistance in sewage treatment plants . https://microbiology.se/2016/08/17/published-paper-antibiotic-resistance-in-sewage-treatment-plants/
- Flach C-F, Pal C, Svensson CJ, Kristiansson E, Östman M, Bengtsson-Palme J, Tysklind M, Larsson DGJ: Does antifouling paint select for antibiotic resistance? Science of the Total Environment, in press (2017). doi: 10.1016/j.scitotenv.2017.01.213 [Paper link]
- Pal C, Bengtsson-Palme J, Kristiansson E, Larsson DGJ: Co-occurrence of resistance genes to antibiotics, biocides and metals reveals novel insights into their co-selection potential. BMC Genomics, 16, 964 (2015). doi: 10.1186/s12864-015-2153-5 [Paper link]
Reflections from the NGS Congress 2016
As the 8th Next Generation Sequencing Congress in London is drawing to a close as I write this, I have a few reflections that might warrant sharing. The first thing that has been apparent this year compared to the two previous times I have visited the event (in 2012 and 2013) is that there was very little talk about where Illumina sequencing is heading next. Instead the discussion was about the applications of Illumina sequencing in the clinical setting; so apparently this is now so mainstream that we only expect slow progress towards longer reads. Apart from that, Illumina is a completed, mature technology. Instead, the flashlight is now pointing entirely towards long-read sequencing (PacBio, NanoPore) as the next big thing. However, the excitement around these technologies has also sort of faded compared to in 2013 when they were soon-to-arrive. Indeed, it seems like there’s not much to be excited about in the sequencing field at the moment, or at least Oxford Global (who are hosting the conference) has failed to get these technologies here.
What also strikes me is the vast amounts of talk about RNAseq of cancer cells. The scope of this event has narrowed dramatically in the past three years. Which makes me substantially less interested in returning next year. If there is not much to be excited about, and the focus is only on cancer sequencing – despite the human microbiota being a very hot topic at the moment – what is the reason for non-cancer researchers to come to the event? There will need to be a stark shift towards another direction of this event if the arrangers want it to remain a broad NGS event. Otherwise, they may just as well go all in and rename the event the Next Generation Sequencing of Cancer Congress. But I hope they choose to widen the scope again; conferences discussing technology as a foundation for a variety of applications are important meeting points and spawning grounds for novel ideas.
Published paper: Evaluating taxonomic classification software
Yesterday, Molecular Ecology Resources put online an unedited version of a recent paper which I co-authored. This time, Rodney Richardson at Ohio State University has made a tremendous work of evaluating three taxonomic classification software – the RDP Naïve Bayesian Classifier, RTAX and UTAX – on a set of DNA barcoding regions commonly used for plants, namely the ITS2, and the matK, rbcL, trnL and trnH genes.
In the paper (1), we discuss the results, merits and limitations of the classifiers. In brief, we found that:
- There is a considerable trade-off between accuracy and sensitivity for the classifiers tested, which indicates a need for improved sequence classification tools (2)
- UTAX was superior with respect to error rate, but was exceedingly stringent and thus suffered from a low assignment rate
- The RDP Naïve Bayesian Classifier displayed high sensitivity and low error at the family and order levels, but had a genus-level error rate of 9.6 percent
- RTAX showed high sensitivity at all taxonomic ranks, but at the same time consistently produced the high error rates
- The choice of locus has significant effects on the classification sensitivity of all tested tools
- All classifiers showed strong relationships between database completeness, classification sensitivity and classification accuracy
We believe that the methods of comparison we have used are simple and robust, and thereby provides a methodological and conceptual foundation for future software evaluations. On a personal note, I will thoroughly enjoy working with Rodney and Reed again; I had a great time discussing the ins and outs of taxonomic classification with them! The paper can be found here.
References and notes
- Richardson RT, Bengtsson-Palme J, Johnson RM: Evaluating and Optimizing the Performance of Software Commonly Used for the Taxonomic Classification of DNA Sequence Data. Molecular Ecology Resources, Early view (2016). doi: 10.1111/1755-0998.12628 [Paper link]
- This is something that several classifiers also showed in the evaluation we did for the Metaxa2 paper (3). Interestingly enough, Metaxa2 is better at maintaining high accuracy also as sensitivity is increased.
- Bengtsson-Palme J, Hartmann M, Eriksson KM, Pal C, Thorell K, Larsson DGJ, Nilsson RH: Metaxa2: Improved identification and taxonomic classification of small and large subunit rRNA in metagenomic data. Molecular Ecology Resources, 15, 6, 1403–1414 (2015). doi: 10.1111/1755-0998.12399 [Paper link]
Published paper: The global resistome
Late yesterday, Microbiome put online our most recent work, covering the resistomes to antibiotics, biocides and metals across a vast range of environments. In the paper (1), we perform the largest characterization of resistance genes, mobile genetic elements (MGEs) and bacterial taxonomic compositions to date, covering 864 different metagenomes from humans (350), animals (145) and external environments such as soil, water, sewage, and air (369 in total). All the investigated metagenomes were sequenced to at least 10 million reads each, using Illumina technology, making the results more comparable across environments than in previous studies (2-4).
We found that the environment types had clear differences both in terms of resistance profiles and bacterial community composition. Humans and animals hosted microbial communities with limited taxonomic diversity as well as low abundance and diversity of biocide/metal resistance genes and MGEs. On the contrary, the abundance of ARGs was relatively high in humans and animals. External environments, on the other hand, showed high taxonomic diversity and high diversity of biocide/metal resistance genes and MGEs. Water, sediment and soil generally carried low relative abundance and few varieties of known ARGs, whereas wastewater and sludge were on par with the human gut. The environments with the largest relative abundance and diversity of ARGs, including genes encoding resistance to last resort antibiotics, were those subjected to industrial antibiotic pollution and air samples from a Beijing smog event.
A paper investigating this vast amount of data is of course hard to describe in a blog post, so I strongly suggest the interested reader to head over to Microbiome’s page and read the full paper (1). However, here’s a ver short summary of the findings:
- The median relative abundance of ARGs across all environments was 0.035 copies per bacterial 16S rRNA
- Antibiotic-polluted environments have (by far) the highest abundances of ARGs
- Urban air samples carried high abundance and diversity of ARGs
- Human microbiota has high abundance and diversity of known ARGs, but low taxonomic diversity compared to the external environment
- The human and animal resistomes are dominated by tetracycline resistance genes
- Over half of the ARGs were only detected in external environments, while 20.5 % were found in human, animal and at least one of the external environments
- Biocide and metal resistance genes are more common in external environments than in the human microbiota
- Human microbiota carries low abundance and richness of MGEs compared to most external environments
Importantly, less than 1.5 % of all detected ARGs were found exclusively in the human microbiome. At the same time, 57.5 % of the known ARGs were only detected in metagenomes from environmental samples, despite that the majority of the investigated ARGs were initially encountered in pathogens. Still, our analysis suggests that most of these genes are relatively rare in the human microbiota. Environmental samples generally contained a wider distribution of resistance genes to a more diverse set of antibiotics classes. For example, the relative abundance of beta-lactam resistance genes was much larger in external environments than in human and animal microbiomes. This suggests that the external environment harbours many more varieties of resistance genes than the ones currently known from the clinic. Indeed, functional metagenomics has resulted in the discovery of many novel ARGs in external environments (e.g. 5). This all fits well with an overall much higher taxonomic diversity of environmental microbial communities. In terms of consequences associated with the potential transfer of ARGs to human pathogens, we argue that unknown resistance genes are of greater concern than those already known to circulate among human-associated bacteria (6).
This study describes the potential for many external environments, including those subjected to pharmaceutical pollution, air and wastewater/sludge, to serve as hotspots for resistance development and/or transmission of ARGs. In addition, our results indicate that these environments may play important roles in the mobilization of yet unknown ARGs and their further transmission to human pathogens. To provide guidance for risk-reducing actions we – based on this study – suggest strict regulatory measures of waste discharges from pharmaceutical industries and encourage more attention to air in the transmission of antibiotic resistance (1).
References
- Pal C, Bengtsson-Palme J, Kristiansson E, Larsson DGJ: The structure and diversity of human, animal and environmental resistomes. Microbiome, 4, 54 (2016). doi: 10.1186/s40168-016-0199-5
- Durso LM, Miller DN, Wienhold BJ. Distribution and quantification of antibiotic resistant genes and bacteria across agricultural and non-agricultural metagenomes. PLoS One. 2012;7:e48325.
- Nesme J, Delmont TO, Monier J, Vogel TM. Large-scale metagenomic-based study of antibiotic resistance in the environment. Curr Biol. 2014;24:1096–100.
- Fitzpatrick D, Walsh F. Antibiotic resistance genes across a wide variety of metagenomes. FEMS Microbiol Ecol. 2016. doi:10.1093/femsec/fiv168.
- Allen HK, Moe LA, Rodbumrer J, Gaarder A, Handelsman J. Functional metagenomics reveals diverse β-lactamases in a remote Alaskan soil. ISME J. 2009;3:243–51.
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
Database quality paper in special issue
I just want to highlight that the paper on strategies to improve database accuracy and usability we recently published in Proteomics (1) has been included in their most recent issue, which is a special issue focusing on Data Quality Issues in Proteomics. I highly recommend reading our paper (of course) and many of the other in the special issue. Happy reading!
On another note, I will be giving a talk next Wednesday (October 5th) on a seminar day on next generation sequencing in clinical microbiology, titled “Antibiotic resistance in the clinic and the environment – There and back again“. You are very welcome to the lecture hall at floor 3 in our building at Guldhedsgatan 10A here in Gothenburg if you are interested! (Bear in mind though that it all starts at 8.15 in the morning.)
Finally, it seems that I am going to the Next Generation Sequencing Congress in London this year, which will be very fun! Hope to see some of you dealing with sequencing there!
References
- Bengtsson-Palme J, Boulund F, Edström R, Feizi A, Johnning A, Jonsson VA, Karlsson FH, Pal C, Pereira MB, Rehammar A, Sánchez J, Sanli K, Thorell K: Strategies to improve usability and preserve accuracy in biological sequence databases. Proteomics, 16, 18, 2454–2460 (2016). doi: 10.1002/pmic.201600034 [Paper link]
Published paper: Annotating fungi from the built environment
MycoKeys today put a paper online which I was involved in. The paper describes the results of a workshop in May, when we added and refined annotations for fungal ITS sequences according to the MIxS-Built Environment annotation standard (1). Fungi have been associated with a range of unwanted effects in the built environment, including asthma, decay of building materials, and food spoilage. However, the state of the metadata annotation of fungal DNA sequences from the built environment is very much incomplete in public databases. The workshop aimed to ease a little part of this problem, by distributing the re-annotation of public fungal ITS sequences across 36 persons. In total, we added or changed of 45,488 data points drawing from published literature, including addition of 8,430 instances of countries of collection, 5,801 instances of building types, and 3,876 instances of surface-air contaminants. The results have been implemented in the UNITE database and shared with other online resources. I believe, that distributed initiatives like this (and the ones I have been involved in in the past (2,3)) serve a very important purpose for establishing better annotation of sequence data, an issue I have brought up also for sequences outside of barcoding genes (4). The full paper can be found here.
References
- Abarenkov K, Adams RI, Laszlo I, Agan A, Ambrioso E, Antonelli A, Bahram M, Bengtsson-Palme J, Bok G, Cangren P, Coimbra V, Coleine C, Gustafsson C, He J, Hofmann T, Kristiansson E, Larsson E, Larsson T, Liu Y, Martinsson S, Meyer W, Panova M, Pombubpa N, Ritter C, Ryberg M, Svantesson S, Scharn R, Svensson O, Töpel M, Untersehrer M, Visagie C, Wurzbacher C, Taylor AFS, Kõljalg U, Schriml L, Nilsson RH: Annotating public fungal ITS sequences from the built environment according to the MIxS-Built Environment standard – a report from a May 23-24, 2016 workshop (Gothenburg, Sweden). MycoKeys, 16, 1–15 (2016). doi: 10.3897/mycokeys.16.10000
- Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M, Bates ST, Bruns TT, Bengtsson-Palme J, Callaghan TM, Douglas B, Drenkhan T, Eberhardt U, Dueñas M, Grebenc T, Griffith GW, Hartmann M, Kirk PM, Kohout P, Larsson E, Lindahl BD, Lücking R, Martín MP, Matheny PB, Nguyen NH, Niskanen T, Oja J, Peay KG, Peintner U, Peterson M, Põldmaa K, Saag L, Saar I, Schüßler A, Senés C, Smith ME, Suija A, Taylor DE, Telleria MT, Weiß M, Larsson KH: Towards a unified paradigm for sequence-based identification of Fungi. Molecular Ecology, 22, 21, 5271–5277 (2013). doi: 10.1111/mec.12481
- Nilsson RH, Hyde KD, Pawlowska J, Ryberg M, Tedersoo L, Aas AB, Alias SA, Alves A, Anderson CL, Antonelli A, Arnold AE, Bahnmann B, Bahram M, Bengtsson-Palme J, Berlin A, Branco S, Chomnunti P, Dissanayake A, Drenkhan R, Friberg H, Frøslev TG, Halwachs B, Hartmann M, Henricot B, Jayawardena R, Jumpponen A, Kauserud H, Koskela S, Kulik T, Liimatainen K, Lindahl B, Lindner D, Liu J-K, Maharachchikumbura S, Manamgoda D, Martinsson S, Neves MA, Niskanen T, Nylinder S, Pereira OL, Pinho DB, Porter TM, Queloz V, Riit T, Sanchez-García M, de Sousa F, Stefaczyk E, Tadych M, Takamatsu S, Tian Q, Udayanga D, Unterseher M, Wang Z, Wikee S, Yan J, Larsson E, Larsson K-H, Kõljalg U, Abarenkov K: Improving ITS sequence data for identification of plant pathogenic fungi. Fungal Diversity, 67, 1, 11–19 (2014). doi: 10.1007/s13225-014-0291-8
- Bengtsson-Palme J, Boulund F, Edström R, Feizi A, Johnning A, Jonsson VA, Karlsson FH, Pal C, Pereira MB, Rehammar A, Sánchez J, Sanli K, Thorell K: Strategies to improve usability and preserve accuracy in biological sequence databases. Proteomics, Early view (2016). doi: 10.1002/pmic.201600034