The Lennart Sparell Prize
I am happy to announce that Cancer- och Allergifonden [the Cancer and Allergy Foundation] have awarded me with the first Lennart Sparell prize. The prize was instated in memory of the foundations founder – Lennart Sparell, who passed away last year – and is awarded to researchers (or other persons) who have thought outside-of-the-box or challenged the current paradigms. A particular emphasis is given to research on environmental pollutants that affect human health through food or environmental exposure.
Naturally, I am honored to be the recipient of this prize. The award was motivated by the research I have done on the role of ecological and evolutionary processes in the external environment in driving antibiotic resistance development, and how that can have consequences for human health. Particularly, I am happy that the research that I, Joakim Larsson, Erik Kristiansson and a few others on the role of environmental processes in the development of antibiotic resistance and the recruitment of novel resistance genes are given attention. This view, which perhaps do not challenge the paradigm but at the very least points to an alternative risk scenario, has often been neglected when environmental antibiotic resistance has been discussed.
The prize will be awarded on a ceremony on June 10 in Stockholm, but I would already now take the opportunity to thank everyone who has been involved in the research being recognized, particularly Joakim Larsson and Erik Kristiansson – this award is to a very very large extent to your merit.
Published paper: A novel Na-binding site in sialic acid symporters
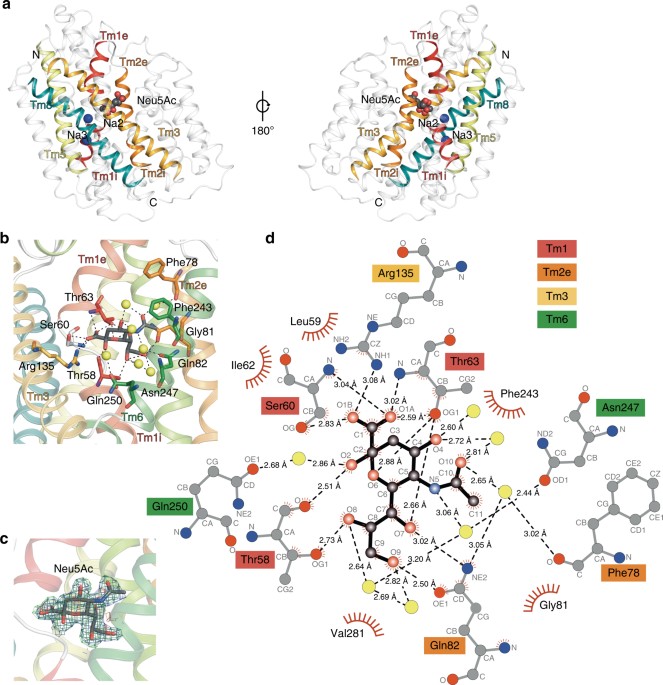
I have been quite occupied with other things the last couple of days, so I am late on the ball here. Anyway, on May 1st, Nature Communications published a paper on the protein structure of SiaT, a sialic acid transporter from Proteus mirabilis (1). Many pathogens use sialic acids as an energy source or as an external coating to evade the immune defense (2). Therefore, many bacteria that colonize sialylated environments have transporters which specifically import sialic acids. SiaT is one of those transporters, belonging to the sodium solute symporter (SSS) family (3) (with for some weird reason is associated with the Pfam family “SSF”, an eternal source of confusion in discussions within this project). The SSS proteins use Na+ gradients to drive the import of desired substrates (4). Based on the protein structure, our team found that SiaT binds two Na+ ions. One binds to the conserved, well-known, Na2 site, but the other Na+ binds to a new position, which we term Na3. This position (this is where my part of the work comes in) is conserved in many SSS family members. We finally used functional and molecular dynamics studies to validate the substrate-binding site and demonstrate that both Na+ sites regulate N-acetylneuraminic acid transport.
As I hinted, i am not venturing into protein structures – that part of this work has been performed by an excellent team associated with Dr. Rosmarie Friemann. Instead, my part is essentially summarized in these two sentences of the manuscript: “We analysed all SSS sequences that contained the primary Na2 site (21,467) to determine the degree of conservation of the Na3 site, allowing for threonine at either Ser345 or Ser346. Na3 is present in 19.6% (4212) of these sequences including hSGLT1, which transports two Na+, but not vSGLT or hSGLT2, which transport only one Na+” (1). That’s a few months of works condensed into 55 words. Still, the exciting thing here is that we find an evolutionary conserved Na-binding site, which has so far eluded detection.
The results of this work provides a better understanding of how secondary active transporters harness additional energy from ion gradients. It may be possible to exploit differences in this mechanism between different SSS family members (and other transporters with the LeuT fold) to develop new antimicrobials, something that is urgently needed in the face of the rapidly increasing antibiotic resistance.
References
- Wahlgren WY°, North RA°, Dunevall E°, Paz A, Scalise M, Bisognano P, Bengtsson-Palme J, Goyal P, Claesson E, Caing-Carlsson R, Andersson R, Beis K, Nilsson U, Farewell A, Pochini L, Indiveri C, Grabe M, Dobson RCJ, Abramson J, Ramaswamy S, Friemann R: Substrate-bound outward-open structure of a Na+-coupled sialic acid symporter reveals a novel Na+ site. Nature Communications, 9, 1753 (2018). doi: 10.1038/s41467-018-04045-7
- Vimr ER, Kalivoda KA, Deszo EL, Steenburgen SM: Diversity of microbial sialic acid metabolism. Microbiology and Molecular Biology Reviews, 68, 132–153 (2004).
- North RA, Horne CR, Davies JS, Remus DM, Muscroft-Taylor AC, Goyal P, Wahlgren WY, Ramaswamy S, Friemann R, Dobson RCJ: “Just a spoonful of sugar…”: import of sialic acid across bacterial cell membranes. Biophysical Reviews, 10, 219–227 (2017).
- Severi E, Hosie AH, Hawkhead JA, Thomas GH: Characterization of a novel sialic acid transporter of the sodium solute symporter (SSS) family and in vivo comparison with known bacterial sialic acid transporters. FEMS Microbiology Letters, 304, 47–54 (2010).
Published paper: Environmental factors leading to resistance
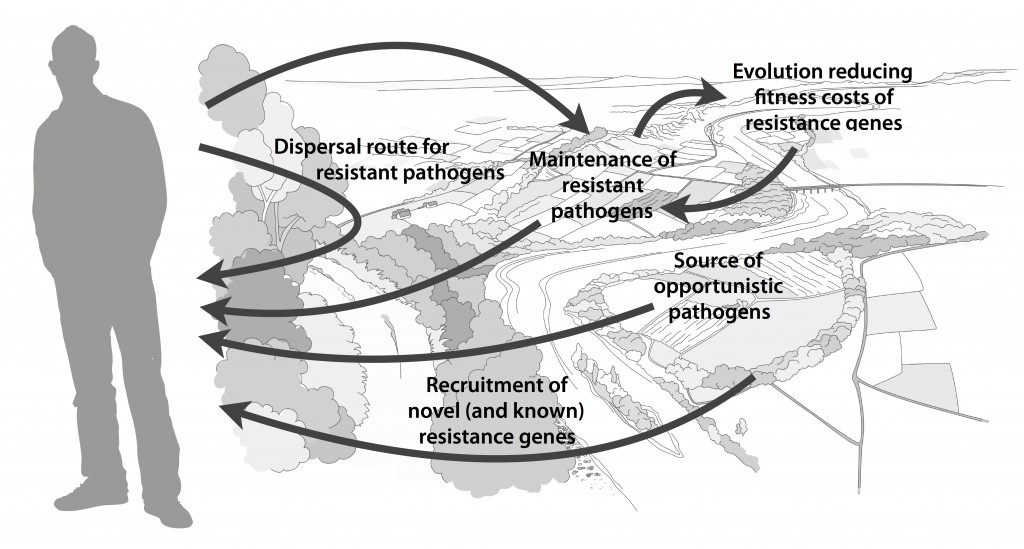
Myself, Joakim Larsson and Erik Kristiansson have written a review on the environmental factors that influence development and spread of antibiotic resistance, which was published today in FEMS Microbiology Reviews. The review (1) builds on thoughts developed in the latter parts of my PhD thesis (2), and seeks to provide a synthesis knowledge gained from different subfields towards the current understanding of evolutionary and ecological processes leading to clinical appearance of resistance genes, as well as the important environmental dispersal barriers preventing spread of resistant pathogens.
We postulate that emergence of novel resistance factors and mobilization of resistance genes are likely to occur continuously in the environment. However, the great majority of such genetic events are unlikely to lead to establishment of novel resistance factors in bacterial populations, unless there is a selection pressure for maintaining them or their fitness costs are negligible. To enable measures to prevent resistance development in the environment, it is therefore critical to investigate under what conditions and to what extent environmental selection for resistance takes place. Selection for resistance is likely less important for the dissemination of resistant bacteria, but will ultimately depend on how well the species or strain in question thrives in the external environment. Metacommunity theory (3,4) suggests that dispersal ability is central to this process, and therefore opportunistic pathogens with their main habitat in the environment may play an important role in the exchange of resistance factors between humans and the environment. Understanding the dispersal barriers hindering this exchange is not only key to evaluate risks, but also to prevent resistant pathogens, as well as novel resistance genes, from reaching humans.
Towards the end of the paper, we suggest certain environments that seem to be more important from a risk management perspective. We also discuss additional problems linked to the development of antibiotic resistance, such as increased evolvability of bacterial genomes (5) and which other types of genes that may be mobilized in the future, should the development continue (1,6). In this review, we also further develop thoughts on the relative risks of re-recruiting and spreading well-known resistance factors already circulating in pathogens, versus recruitment of completely novel resistance genes from environmental bacteria (7). While the latter case is likely to be very rare, and thus almost impossible to quantify the risks for, the consequences of such (potentially one-time) events can be dire.
I personally think that this is one of the best though-through pieces I have ever written, and since it is open access and (in my biased opinion) written in a fairly accessible way, I recommend everyone to read it. It builds on the ecological theories for resistance ecology developed by, among others, Fernando Baquero and José Martinez (8-13). Over the last year, it has been stressed several times at meetings (e.g. at the EDAR conferences in August) that there is a need to develop an ecological framework for antibiotic resistance genes. I think this paper could be one of the foundational pillars on such an endeavor and look forward to see how it will fit into the growing literature on the subject!
References
- Bengtsson-Palme J, Kristiansson E, Larsson DGJ: Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews, accepted manuscript (2017). doi: 10.1093/femsre/fux053
- Bengtsson-Palme J: Antibiotic resistance in the environment: a contribution from metagenomic studies. Doctoral thesis (medicine), Department of Infectious Diseases, Institute of Biomedicine, Sahlgrenska Academy, University of Gothenburg, 2016. [Link]
- Bengtsson J: Applied (meta)community ecology: diversity and ecosystem services at the intersection of local and regional processes. In: Verhoef HA, Morin PJ (eds.). Community Ecology: Processes, Models, and Applications. Oxford: Oxford University Press, 115–130 (2009).
- Leibold M, Norberg J: Biodiversity in metacommunities: Plankton as complex adaptive systems? Limnology and Oceanography, 1278–1289 (2004).
- Gillings MR, Stokes HW: Are humans increasing bacterial evolvability? Trends in Ecology and Evolution, 27, 346–352 (2012).
- Gillings MR: Evolutionary consequences of antibiotic use for the resistome, mobilome and microbial pangenome. Frontiers in Microbiology, 4, 4 (2013).
- Bengtsson-Palme J, Larsson DGJ: Antibiotic resistance genes in the environment: prioritizing risks. Nature Reviews Microbiology, 13, 369 (2015). doi: 10.1038/nrmicro3399-c1
- Baquero F, Alvarez-Ortega C, Martinez JL: Ecology and evolution of antibiotic resistance. Environmental Microbiology Reports, 1, 469–476 (2009).
- Baquero F, Tedim AP, Coque TM: Antibiotic resistance shaping multi-level population biology of bacteria. Frontiers in Microbiology, 4, 15 (2013).
- Berendonk TU, Manaia CM, Merlin C et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015).
- Hiltunen T, Virta M, Laine A-L: Antibiotic resistance in the wild: an eco-evolutionary perspective. Philosophical Transactions of the Royal Society B: Biological Sciences, 372 (2017) doi: 10.1098/rstb.2016.0039.
- Martinez JL: Bottlenecks in the transferability of antibiotic resistance from natural ecosystems to human bacterial pathogens. Frontiers in Microbiology, 2, 265 (2011).
- Salyers AA, Amábile-Cuevas CF: Why are antibiotic resistance genes so resistant to elimination? Antimicrobial Agents and Chemotherapy, 41, 2321–2325 (1997).
Published paper: Antibiotic resistance in sewage treatment plants
 After a long wait (1), Science of the Total Environment has finally decided to make our paper on selection of antibiotic resistance genes in sewage treatment plants (STPs) available (2). STPs are often suggested to be “hotspots” for emergence and dissemination of antibiotic-resistant bacteria (3-6). However, we actually do not know if the selection pressures within STPs, that can be caused either by residual antibiotics or other co-selective agents, are sufficiently large to specifically promote resistance. To better understand this, we used shotgun metagenomic sequencing of samples from different steps of the treatment process (incoming water, treated water, primary sludge, recirculated sludge and digested sludge) in three Swedish STPs in the Stockholm area to characterize the frequencies of resistance genes to antibiotics, biocides and metal, as well as mobile genetic elements and taxonomic composition. In parallel, we also measured concentrations of antibiotics, biocides and metals.
After a long wait (1), Science of the Total Environment has finally decided to make our paper on selection of antibiotic resistance genes in sewage treatment plants (STPs) available (2). STPs are often suggested to be “hotspots” for emergence and dissemination of antibiotic-resistant bacteria (3-6). However, we actually do not know if the selection pressures within STPs, that can be caused either by residual antibiotics or other co-selective agents, are sufficiently large to specifically promote resistance. To better understand this, we used shotgun metagenomic sequencing of samples from different steps of the treatment process (incoming water, treated water, primary sludge, recirculated sludge and digested sludge) in three Swedish STPs in the Stockholm area to characterize the frequencies of resistance genes to antibiotics, biocides and metal, as well as mobile genetic elements and taxonomic composition. In parallel, we also measured concentrations of antibiotics, biocides and metals.
We found that only the concentrations of tetracycline and ciprofloxacin in the influent water were above those that we predict to cause resistance selection (7). However, there was no consistent enrichment of resistance genes to any particular class of antibiotics in the STPs, neither for biocide and metal resistance genes. Instead, the most substantial change of the bacterial communities compared to human feces (sampled from Swedes in another study of ours (8)) occurred already in the sewage pipes, and was manifested by a strong shift from obligate to facultative anaerobes. Through the treatment process, resistance genes against antibiotics, biocides and metals were not reduced to the same extent as fecal bacteria were.
Worryingly, the OXA-48 beta-lactamase gene was consistently enriched in surplus and digested sludge. OXA-48 is still rare in Swedish clinical isolates (9), but provides resistance to carbapenems, one of our most critically important classes of antibiotics. However, taken together metagenomic sequencing did not provide clear support for any specific selection of antibiotic resistance. Rather, since stronger selective forces affect gross taxonomic composition, and thereby also resistance gene abundances, it is very hard to interpret the metagenomic data from a risk-for-selection perspective. We therefore think that comprehensive analyses of resistant vs. non-resistant strains within relevant species are warranted.
Taken together, the main take-home messages of the paper (2) are:
- There were no apparent evidence for direct selection of resistance genes by antibiotics or co-selection by biocides or metals
- Abiotic factors (mostly oxygen availability) strongly shape taxonomy and seems to be driving changes of resistance genes
- Metagenomic and/or PCR-based community studies may not be sufficiently sensitive to detect selection effects, as important shifts towards resistant may occur within species and not on the community level
- The concentrations of antibiotics, biocides and metals were overall reduced, but not removed in STPs. Incoming concentrations of antibiotics in Swedish STPs are generally low
- Resistance genes are overall reduced through the treatment process, but far from eliminated
References and notes
- Okay, those who takes notes know that I have already complained once before on Science of the Total Environment’s ridiculously long production handling times. But, seriously, how can a journal’s production team return the proofs for after three days of acceptance, and then wait seven weeks before putting the final proofs online? I still wonder what is going on beyond the scenes, which is totally obscure because the production office also refuses to respond to e-mails. Not a nice publishing experience this time either.
- Bengtsson-Palme J, Hammarén R, Pal C, Östman M, Björlenius B, Flach C-F, Kristiansson E, Fick J, Tysklind M, Larsson DGJ: Elucidating selection processes for antibiotic resistance in sewage treatment plants using metagenomics. Science of the Total Environment, in press (2016). doi: 10.1016/j.scitotenv.2016.06.228 [Paper link]
- Rizzo L, Manaia C, Merlin C, Schwartz T, Dagot C, Ploy MC, Michael I, Fatta-Kassinos D: Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Science of the Total Environment, 447, 345–360 (2013). doi: 10.1016/j.scitotenv.2013.01.032
- Laht M, Karkman A, Voolaid V, Ritz C, Tenson T, Virta M, Kisand V: Abundances of Tetracycline, Sulphonamide and Beta-Lactam Antibiotic Resistance Genes in Conventional Wastewater Treatment Plants (WWTPs) with Different Waste Load. PLoS ONE, 9, e103705 (2014). doi: 10.1371/journal.pone.0103705
- Yang Y, Li B, Zou S, Fang HHP, Zhang T: Fate of antibiotic resistance genes in sewage treatment plant revealed by metagenomic approach. Water Research, 62, 97–106 (2014). doi: 10.1016/j.watres.2014.05.019
- Berendonk TU, Manaia CM, Merlin C, Fatta-Kassinos D, Cytryn E, Walsh F, et al.: Tackling antibiotic resistance: the environmental framework. Nature Reviews Microbiology, 13, 310–317 (2015). doi: 10.1038/nrmicro3439
- Bengtsson-Palme J, Larsson DGJ: Concentrations of antibiotics predicted to select for resistant bacteria: Proposed limits for environmental regulation. Environment International, 86, 140–149 (2016). doi: 10.1016/j.envint.2015.10.015
- Bengtsson-Palme J, Angelin M, Huss M, Kjellqvist S, Kristiansson E, Palmgren H, Larsson DGJ, Johansson A: The human gut microbiome as a transporter of antibiotic resistance genes between continents. Antimicrobial Agents and Chemotherapy, 59, 10, 6551–6560 (2015). doi: 10.1128/AAC.00933-15
- Hellman J, Aspevall O, Bengtsson B, Pringle M: SWEDRES-SVARM 2014. Consumption of antimicrobials and occurrence of antimicrobial resistance in Sweden. Public Health Agency of Sweden and National Veterinary Institute, Solna/Uppsala, Sweden. Report No.: 14027. Available from: http://www.folkhalsomyndigheten.se/publicerat-material/ (2014)
Evolution vs. Succession – what are we really studying?
One thing that I find slightly annoying is when people do not get the basic concepts right – or when debatable concepts are used without discussion of their implications. This further annoys me when it is done by senior scientists, who should know better. Sometimes, I guess this happens out of ignorance, and sometimes to be able to stick your subject to a certain buzzword concept. Neither is good, even though the former reason is little more forgivable then the latter. One area where this problem becomes agonizingly evident is when molecular biologists or medical scientists moves into ecology, as has happened with the advent of metagenomics. When the study of the human gut microflora turned into a large-scale sequencing effort, people who had previously studied bacteria grown on plates started facing a world of community ecology. However, I get the impression that way too often these people do not ask ecologists for advice, or even read up on the ecological literature. Which, I suppose, is the reason why medical scientists can talk about how the human gut microflora can “evolve” into a stable community a couple years after birth, even though words such as “development” or “succession” would be much more accurate to describe this change.
The marker gene flaw
To set what I mean straight, let us compare the human gut to a forest. If an open field is left to itself, larger plants will slowly inhabit it, and gradually different species will replace each other, until we have a fully developed forest. Similarly, the human gut microflora is at birth rather unstable, but stabilizes relatively quickly and within a few years we have a microbial community with “adult-like” characteristics. To arrive at this conclusion, scientists generally use the 16S (small sub-unit) genetic marker to study the bacterial species diversity. This works in pretty much the same way as going out into the forest and count trees of different kinds.
Now, if I went out into the forest once and counted the tree species, waited for 50 years and then did the same thing again, I would presumably see that the forest species composition had changed. However, if I called this “evolution”, fellow scientists would laugh at me. Raspberry bushes do not evolve into birches, and birches do not evolve into firs. Instead, ecologists talk about “succession”; a progressive transformation of a community, going on until a stable community is formed. The concept of succession seems well-suited also to describe what is happening in the human gut, and should of course also be used in that setting. The most likely driver of the functional community changes is not that some bacterial species have evolved new functions, but rather that bacterial species performing these new functions have outcompeted the once previously present.
In fact, I would argue that it is impossible to study evolution through a genetic marker such as the 16S gene (except in the rare case when you study evolution of the 16S gene itself). Instead, the only thing we could assess using a marker gene is how the copy number of the different gene variants change over time (or space, or conditions). The copy number tells us about the species composition of the community at a given time, which can be used to measure successional changes. However, evolutionary changes would require heritable changes in the characteristics of biological populations, i.e. that their genetic material change in some way. Unless that change happens in the marker gene of choice, we cannot measure it, and the alterations of composition we measure will only reflect differences in species abundances. These differences might have arisen from genetic (i.e. evolutionary) changes, but we cannot assess that.
What are we studying with metagenomics?
This brings us to the next problem, which is not only a problem of semantics and me getting annoyed, but a problem with real implications. What are we really studying using metagenomics? When we apply an environmental sequencing approach to a microbial community, we get a snapshot of the genetic material at a given time and site; at specific conditions. Usually, we aim to characterize the community from a taxonomic or functional perspective, and we often have some other community which we want to compare to. However, if we only collect data from different communities at one time point, or if we only study a community before and after exposure, we have no way of telling if differences stem from selective pressures or from more a random succession progress. As most microbial habitats are not as well studied as the human gut, we know little about microbial community assembly and succession.
Also, in ecology a disturbance to a particular community is generally considered as a starting point for a new succession process. This process may, or may not, return the community to the same stable state. However, if the disturbance was of permanent nature, the new community will have to adapt to the new conditions, and the stable state will likely not have the same species distribution. Such an adaption could be caused by genetic changes (which would clearly be an evolutionary process), or by simple replacement of sensitive species with tolerant ones. The latter would be a selective process, but not necessarily an evolutionary one. If the selection does not alter the genetic material within the species, but only the species composition, I would argue that this is also a case of succession.
Complications with resistance
This complicates the work with metagenomic data. If we study antibiotic resistance genes, and say that bacteria in an environment have evolved antibiotic resistance, we base that assertion on that genes responsible for resistance have either evolved within the present bacteria, or have (more likely) been transferred into the genomes of the bacteria via horizontal gene transfer. However, if the resistance profile we see is simply caused by a replacement of sensitive species with resistant ones, we have not really discovered something new evolving, but are only witnessing spread of already resistant bacteria. In the gut, this would be a problem by itself, but say that we do the same study in the open environment. We already know that environmental bacteria have contained resistance genes for ages, so the real threat to human health here would be a spread from naturally resistant bacteria to human pathogens. However, as mentioned earlier, without extremely well thought-through methodology we cannot really see such transmissions of resistance genes. Here, the search for mobile elements, and large-scale takes on community composition vs. resistance profiles in contaminated and non-polluted areas can play a huge role in shedding light on the question of spreading. However, this will require larger and better planned experiments using metagenomics than what is generally performed at the moment. The questions of microbial community assembly, dispersal, succession and adaption are still largely unanswered, and our metagenomic and environmental sequencing approaches have just started to tinker around with the lid of the jar.